CAZypedia needs your help! We have many unassigned GH, PL, CE, AA, GT, and CBM pages in need of Authors and Responsible Curators.
Scientists at all career stages, including students, are welcome to contribute to CAZypedia. Read more here, and in the 10th anniversary article in Glycobiology.
New to the CAZy classification? Read this first.
*
Consider attending the 15th Carbohydrate Bioengineering Meeting in Ghent, 5-8 May 2024.
Carbohydrate Binding Module Family 29
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
- Author: ^^^Harry Gilbert^^^
- Responsible Curator: ^^^Elizabeth Ficko-Blean^^^
| CAZy DB link | |
| http://www.cazy.org/CBM29.html |
Ligand specificities
The two family 29 CBMs (CBM29-1 and CBM29-2) are located in the same Pyromyces equi protein defined as noncatalytic protein 1 or NCP1 [1]. There are no other members of this family in the CAZy database. NCP1 binds to glucomannans, galactomannans, β-glucans that contain β1,4-linkages, and insoluble forms of mannan and cellulose. The two individual CBM29s in NCP1 bind to all these polysaccharides but with affinities (e.g. glucomannan KA of 37%-1 for CBM29-1 and 228%-1 for CBM29-2) substantially lower than the full length protein (glucomannan KA of 11990%-1), providing a classical example of avidity effects. Ligand binding is enthalpically driven with changes in entropy having a negative impact on affinity, typical of CBMs that bind to soluble ligands. The targeting of soluble polysacchride chains with an endo-mode of binding (individual CBM29s bind to mannose based polysaccharides with a coverage of one protein molecule for 8-9 sugars [2]) classify these proteins as type B CBMs.
Structural Features
[[
File:CBM29E75Rmutant.png|thumb|300px|right|Figure 2. The structure of the E78R mutant of the C-terminal CBM29 of P. equi protein NCP1 in complex with cellohexaose (PDB ID 1OH3).The figure highlights the aromatic residues from the two molecules of the CBM29 that collectively interact with all the pyranose rings of the ligand explaining why the mutant binds to cellohexaose and not mannohexaose, which contains an axial OH at C2 that would make steric clashes with the aromatic residues.]]
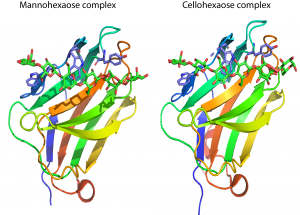
CBM29s comprise ~150 amino acids. The crystal structure of the C-terminal CBM29 module (CBM29-2) from P. equi NCP1 was determined in complex with cellohexaose and mannohexaose [2]. The structure presents a classic β-jelly-roll, with five β-strands forming each of the two faces typical of many other CBM families. [3]. The concave surface of CBM29-2 comprises the ligand binding cleft optimized to bind single glycan chains consistent with its type B designation. In complexes with cellohexaose or mannohexaose the hexasaccharides occupy the binding cleft. Three aromatic residues that line this cleft forms a classic hydrophobic platform observed for many CBMs. All three aromatic amino acids interact with the α-face of the glucosyl or mannosyl moieties of residues 6, 4, and 2, respectively (Figure 1). Discrete recognition of the O3 and O6 hydroxyls builds upon this template, conferring further specificity toward either glucose or mannose-based ligands. Interaction with the 2-hydroxyl, whose chirality defines whether the pyranoside is glucose or mannose, is flexible, thus allowing the relaxed recognition of both gluco- and manno-configured ligands. The mechanism by which CBM16 modules recognize manno- and gluco-configured ligands [4] strongly resembles the structural basis by which CBM29-2 binds to cellohexaose and mannohexaose.
Mutagenesis studies of CBM29-2 reinforce the major role played by the aromatic residues in the binding cleft in ligand recognition, and clarify the importance of the amino acids that are in hydrogen bonding distance with the two ligands [5]. One of the mutants generated, Glu to Arg substitution, altered ligand specificity; the mutant bound to gluco- but not manno-configured ligands. Isothermal titration calorimetry in combination with gel filtration and X-ray crystallography showed that two molecules of the mutant CBM bound to a single cellohexaose molecule to form a trimolecular sandwich complex in solution. Each of the pyranose rings of cellohexaose was interacting with one of the mutant proteins (Figure 2), explaining how the axial O2 of mannose prevented ligand binding [6]. A similar trimolecular complex in crystal, comprising two molecules of a CBM63 expansin that sandwiches a cellohexaose molecule, has also been described [7].
Functionalities
The single non-catalytic protein NCP1 contains three N-terminal dockerin domains (Glu22 to Arg147) and two CBM29 modules extending form Arg172 to Val327 (CBM29-1) and Lys332 to Asn478 (CBM29-2). The dockerin domains enable NCP1 to be incorporated into the P. equi cellulosome, a multienzyme plant cell wall degrading complex [1, 8]. It was proposed that the plasticity in ligand recognition enabled NCP1 to make a significant contribution to the attachment of the fungal cellulosome to the plant cell wall [1].
Family Firsts
- First Identified
- The pair of CBM29s were first observed in the P. equi cellulosomal protein NCP1 by [1]
- First structural characterization
- The first crystal structure of this family is the C-terminal CBM29 (CBM29-2) from the P. equi non-catalytic protein NCP1 [2].
References
- Freelove AC, Bolam DN, White P, Hazlewood GP, and Gilbert HJ. (2001). A novel carbohydrate-binding protein is a component of the plant cell wall-degrading complex of Piromyces equi. J Biol Chem. 2001;276(46):43010-7. DOI:10.1074/jbc.M107143200 |
- Charnock SJ, Bolam DN, Nurizzo D, Szabó L, McKie VA, Gilbert HJ, and Davies GJ. (2002). Promiscuity in ligand-binding: The three-dimensional structure of a Piromyces carbohydrate-binding module, CBM29-2, in complex with cello- and mannohexaose. Proc Natl Acad Sci U S A. 2002;99(22):14077-82. DOI:10.1073/pnas.212516199 |
- Boraston AB, Bolam DN, Gilbert HJ, and Davies GJ. (2004). Carbohydrate-binding modules: fine-tuning polysaccharide recognition. Biochem J. 2004;382(Pt 3):769-81. DOI:10.1042/BJ20040892 |
- Bae B, Ohene-Adjei S, Kocherginskaya S, Mackie RI, Spies MA, Cann IK, and Nair SK. (2008). Molecular basis for the selectivity and specificity of ligand recognition by the family 16 carbohydrate-binding modules from Thermoanaerobacterium polysaccharolyticum ManA. J Biol Chem. 2008;283(18):12415-25. DOI:10.1074/jbc.M706513200 |
- Flint J, Bolam DN, Nurizzo D, Taylor EJ, Williamson MP, Walters C, Davies GJ, and Gilbert HJ. (2005). Probing the mechanism of ligand recognition in family 29 carbohydrate-binding modules. J Biol Chem. 2005;280(25):23718-26. DOI:10.1074/jbc.M501551200 |
- Flint J, Nurizzo D, Harding SE, Longman E, Davies GJ, Gilbert HJ, and Bolam DN. (2004). Ligand-mediated dimerization of a carbohydrate-binding molecule reveals a novel mechanism for protein-carbohydrate recognition. J Mol Biol. 2004;337(2):417-26. DOI:10.1016/j.jmb.2003.12.081 |
- Georgelis N, Yennawar NH, and Cosgrove DJ. (2012). Structural basis for entropy-driven cellulose binding by a type-A cellulose-binding module (CBM) and bacterial expansin. Proc Natl Acad Sci U S A. 2012;109(37):14830-5. DOI:10.1073/pnas.1213200109 |
- Fanutti C, Ponyi T, Black GW, Hazlewood GP, and Gilbert HJ. (1995). The conserved noncatalytic 40-residue sequence in cellulases and hemicellulases from anaerobic fungi functions as a protein docking domain. J Biol Chem. 1995;270(49):29314-22. DOI:10.1074/jbc.270.49.29314 |