CAZypedia needs your help! We have many unassigned GH, PL, CE, AA, GT, and CBM pages in need of Authors and Responsible Curators.
Scientists at all career stages, including students, are welcome to contribute to CAZypedia. Read more here, and in the 10th anniversary article in Glycobiology.
New to the CAZy classification? Read this first.
*
Consider attending the 15th Carbohydrate Bioengineering Meeting in Ghent, 5-8 May 2024.
Difference between revisions of "Carbohydrate Binding Module Family 55"
| Line 18: | Line 18: | ||
== Ligand specificities == | == Ligand specificities == | ||
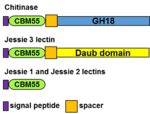
[[File:CBM55.png|thumb|150px|right|'''Figure 1.''' ''Entamoeba'' proteins containing CBM55.]] | [[File:CBM55.png|thumb|150px|right|'''Figure 1.''' ''Entamoeba'' proteins containing CBM55.]] | ||
| − | A CBM55, which is ~60-aa long and contains eight cysteines in conserved positions, was first identified at the N-terminus of the [[GH18]] chitinase of ''Entamoeba histolytica'', the protist that causes dysentery and liver abscess (Figure 1) <cite>Van_Dellen2002,de_la_Vega1997</cite>. CBM55 members are also present at the N-termini of Jessie 3 lectins, which contain a self-aggregating (daub) domain, while CBM55 is the only domain in Jessie 1 and Jessie 2 lectins | + | A CBM55, which is ~60-aa long and contains eight cysteines in conserved positions, was first identified at the N-terminus of the [[GH18]] chitinase of ''Entamoeba histolytica'', the protist that causes dysentery and liver abscess (Figure 1) <cite>Van_Dellen2002,de_la_Vega1997</cite>. CBM55 members are also present at the N-termini of Jessie 3 lectins, which contain a self-aggregating (daub) domain, while CBM55 is the only domain in Jessie 1 and Jessie 2 lectins <cite>Chatterjee2009</cite>. CBM55 members are present in [[GH18]] chitinases and Jessie lectins of all five ''Entamoeba'' species that have been sequenced. CBM55 members are absent from other eukaryotes, eubacteria, and archaea, and so the CBM55 motif appears to have been “created from scratch” by the common ancestor to ''Entamoebae''. CBM55 members of ''E. histolytica'' chitinase, Jessie 1, and Jessie 3 lectins, each produced by gene expression under a constitutive actin promoter in transformed trophozoites, demonstrated binding to chitin beads <cite>Van_Dellen2002</cite>. |
Revision as of 07:42, 22 October 2019
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^John Samuelson^^^
- Responsible Curator: ^^^Elizabeth Ficko-Blean^^^
| CAZy DB link | |
| http://www.cazy.org/CBM55.html |
Ligand specificities
A CBM55, which is ~60-aa long and contains eight cysteines in conserved positions, was first identified at the N-terminus of the GH18 chitinase of Entamoeba histolytica, the protist that causes dysentery and liver abscess (Figure 1) [1, 2]. CBM55 members are also present at the N-termini of Jessie 3 lectins, which contain a self-aggregating (daub) domain, while CBM55 is the only domain in Jessie 1 and Jessie 2 lectins [3]. CBM55 members are present in GH18 chitinases and Jessie lectins of all five Entamoeba species that have been sequenced. CBM55 members are absent from other eukaryotes, eubacteria, and archaea, and so the CBM55 motif appears to have been “created from scratch” by the common ancestor to Entamoebae. CBM55 members of E. histolytica chitinase, Jessie 1, and Jessie 3 lectins, each produced by gene expression under a constitutive actin promoter in transformed trophozoites, demonstrated binding to chitin beads [1].
Structural Features
While the presence of eight cysteines in conserved positions suggest that CBM55 is a disulfide knot, there is no structure for CBM55 [1]. Because CBM55 binds to chitin fibrils in the cyst wall, it is a type A CBM [3].
Functionalities
In the human intestine, motile E. histolytica trophozoites phagocytose bacteria and/or attach to and take bites out of (trogocytose) host epithelial cells [4]. Alternatively, trophozoites form cysts, which have chitin and deacetylated chitin (chitosan) in their wall [5, 6]. E. histolytica is transmitted when cysts are shed in the feces and contaminate food, soil, or water. When cysts are ingested, trophozoites escape their wall and are released into the intestine, completing the life cycle. The most abundant proteins identified by mass spectrometry of cyst walls of E. invadens, a model for the human pathogen, are chitinases, Jessie 3 lectins, and Jacob lectins, which contain two or three chitin-binding domains each with six conserved cysteines [7, 8]. Mass spectrometry and specific antibodies have confirmed the presence of chitinase, Jessie lectins, and Jacob lectins in cyst walls of E. histolytica, the human pathogen [9, 10]. Cyst wall proteins, which are absent from trophozoites but abundant in cysts, coat chitin fibrils and protect them from bacterial chitinases, which are ubiquitous in the environment [11, 12]. One function of CBM55 is to retain chitinase and Jessie 3 lectins in the cyst wall. The chitinase is likely involved in remodeling the cyst wall during encystation and breaking down the cyst wall when the trophozoite escapes during excystation. The daub domain of Jessie 3, which forms a biofilm when produced recombinantly as a fusion with maltose-binding protein in the periplasm of bacteria, makes the cyst wall impenetrable to molecules as small as one kilodalton [3].
Family Firsts
- First Identified
- A CBM55 was first identified at the N-terminus of the GH18 chitinase of Entamoeba histolytica [1, 2].
- First Structural Characterization
- There is no structure available.
References
- Van Dellen K, Ghosh SK, Robbins PW, Loftus B, and Samuelson J. (2002). Entamoeba histolytica lectins contain unique 6-Cys or 8-Cys chitin-binding domains. Infect Immun. 2002;70(6):3259-63. DOI:10.1128/IAI.70.6.3259-3263.2002 |
- de la Vega H, Specht CA, Semino CE, Robbins PW, Eichinger D, Caplivski D, Ghosh S, and Samuelson J. (1997). Cloning and expression of chitinases of Entamoebae. Mol Biochem Parasitol. 1997;85(2):139-47. DOI:10.1016/s0166-6851(96)02817-4 |
- Chatterjee A, Ghosh SK, Jang K, Bullitt E, Moore L, Robbins PW, and Samuelson J. (2009). Evidence for a "wattle and daub" model of the cyst wall of entamoeba. PLoS Pathog. 2009;5(7):e1000498. DOI:10.1371/journal.ppat.1000498 |
- Ralston KS, Solga MD, Mackey-Lawrence NM, Somlata, Bhattacharya A, and Petri WA Jr. (2014). Trogocytosis by Entamoeba histolytica contributes to cell killing and tissue invasion. Nature. 2014;508(7497):526-30. DOI:10.1038/nature13242 |
- Van Dellen KL, Bulik DA, Specht CA, Robbins PW, and Samuelson JC. (2006). Heterologous expression of an Entamoeba histolytica chitin synthase in Saccharomyces cerevisiae. Eukaryot Cell. 2006;5(1):203-6. DOI:10.1128/EC.5.1.203-206.2006 |
- Das S, Van Dellen K, Bulik D, Magnelli P, Cui J, Head J, Robbins PW, and Samuelson J. (2006). The cyst wall of Entamoeba invadens contains chitosan (deacetylated chitin). Mol Biochem Parasitol. 2006;148(1):86-92. DOI:10.1016/j.molbiopara.2006.03.002 |
- Frisardi M, Ghosh SK, Field J, Van Dellen K, Rogers R, Robbins P, and Samuelson J. (2000). The most abundant glycoprotein of amebic cyst walls (Jacob) is a lectin with five Cys-rich, chitin-binding domains. Infect Immun. 2000;68(7):4217-24. DOI:10.1128/IAI.68.7.4217-4224.2000 |
- Van Dellen KL, Chatterjee A, Ratner DM, Magnelli PE, Cipollo JF, Steffen M, Robbins PW, and Samuelson J. (2006). Unique posttranslational modifications of chitin-binding lectins of Entamoeba invadens cyst walls. Eukaryot Cell. 2006;5(5):836-48. DOI:10.1128/EC.5.5.836-848.2006 |
- Ghosh SK, Van Dellen KL, Chatterjee A, Dey T, Haque R, Robbins PW, and Samuelson J. (2010). The Jacob2 lectin of the Entamoeba histolytica cyst wall binds chitin and is polymorphic. PLoS Negl Trop Dis. 2010;4(7):e750. DOI:10.1371/journal.pntd.0000750 |
- Ali IK, Haque R, Siddique A, Kabir M, Sherman NE, Gray SA, Cangelosi GA, and Petri WA Jr. (2012). Proteomic analysis of the cyst stage of Entamoeba histolytica. PLoS Negl Trop Dis. 2012;6(5):e1643. DOI:10.1371/journal.pntd.0001643 |
- Samuelson J and Robbins P. (2011). A simple fibril and lectin model for cyst walls of Entamoeba and perhaps Giardia. Trends Parasitol. 2011;27(1):17-22. DOI:10.1016/j.pt.2010.09.002 |
- Ehrenkaufer GM, Weedall GD, Williams D, Lorenzi HA, Caler E, Hall N, and Singh U. (2013). The genome and transcriptome of the enteric parasite Entamoeba invadens, a model for encystation. Genome Biol. 2013;14(7):R77. DOI:10.1186/gb-2013-14-7-r77 |