CAZypedia needs your help! We have many unassigned GH, PL, CE, AA, GT, and CBM pages in need of Authors and Responsible Curators.
Scientists at all career stages, including students, are welcome to contribute to CAZypedia. Read more here, and in the 10th anniversary article in Glycobiology.
New to the CAZy classification? Read this first.
*
Consider attending the 15th Carbohydrate Bioengineering Meeting in Ghent, 5-8 May 2024.
Difference between revisions of "Carbohydrate Binding Module Family 58"
Harry Brumer (talk | contribs) (various edits, links, etc.) |
|||
| Line 21: | Line 21: | ||
== Ligand specificities == | == Ligand specificities == | ||
| − | + | Only a single CBM58 family member has been characterized, the founding member from the neopullulanase SusG of the human gut symbiont ''Bacteroides thetaiotaomicron'' <cite>Koropatkin2010</cite>. The crystal structure of SusG featuring CBM58 revealed that this domain binds to maltoheptaose as well as acarbose, a pseudotetrasaccharide amylase inhibitor. Isothermal titration calorimetry with maltoheptaose and α-cyclodextrin as well as isotherm depletion assays with insoluble cornstarch further confirmed that CBM58 is a starch-specific CBM <cite>Koropatkin2010</cite>. Based upon these data, the CBM58 family is believed to bind exclusively to α1,4-linked glucan structures in starch. | |
| − | Only a single CBM58 family member has been characterized, the founding member from the neopullulanase SusG of the human gut symbiont ''Bacteroides thetaiotaomicron'' <cite>Koropatkin2010</cite>. The crystal structure of SusG featuring CBM58 revealed that this domain binds to maltoheptaose as well as acarbose, a pseudotetrasaccharide amylase inhibitor. Isothermal titration calorimetry with maltoheptaose and α-cyclodextrin as well as isotherm depletion assays with insoluble cornstarch further confirmed that CBM58 is a starch-specific CBM | ||
== Structural Features == | == Structural Features == | ||
| + | The crystal structure of CBM58 from SusG of ''B. thetaiotaomicron'' displays a canonical β-sandwich fold, with a single binding site accomodated by the loops connecting β3 and β4, β6 and β7, and β7 and β8 <cite>Koropatkin2010</cite>. These loops fold over one of the β-sheets. Because CBM58 recognizes internal regions of the polypeptide chain, it can be classified as a type B CBM <cite>Gilbert2013</cite>. Like many starch-specific CBMs <cite>Machovic2006</cite>, the helical α-glucan is cradled by aromatic stacking interactions with two SusG CBM58 aromatic residues, W287 and W299, as well as hydrogen bonding interactions with K304, N330 and Y260 that aid in positioning the adjacent glucose residues that stack with the tryptophans <cite>Koropatkin2010</cite>. | ||
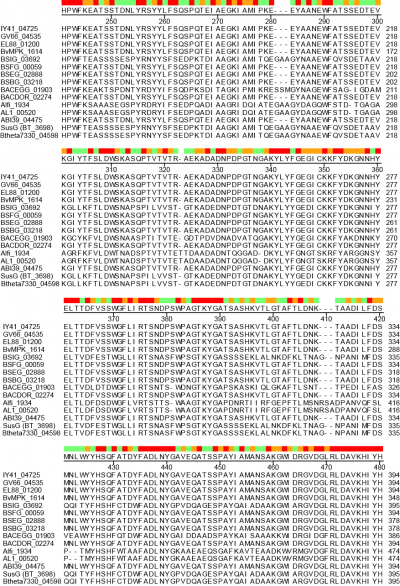
| − | + | A unique facet of the CBM58 of SusG is its position in the middle of the [[GH13]] amylase fold, occupying residues 216-335 of the protein. This domain is inserted between α3 and β3 of the catalytic B domain (residues 153-215 and 336-363), and essentially expands the typically small B domain of GH13 enzymes. Thus the entire SusG sequence can be described from N- to C-terminus (residue numbers): A domain (43-152) - B domain (153-215) - CBM58 (215-335) - B domain (336-363) - A domain (364-607) - C domain (608-692). In SusG, two short linker sequences from residues 212-217 and 334-338, spanning 12Å between the B domain and CBM58, have B-factors and an amino acid sequence that imply this domain is held in a fixed position <cite>Koropatkin2010</cite>. The CBM58 family is small, with the seminal members all arising from the Bacteroidetes phylum. This includes similar [[GH13]] enzymes from isolates of ''Alistipes finegoldii'', ''Alistipes shahii'', ''Bacteroides dorei'', ''Bacteroides eggerthii'' and ''Bacteroides vulgatus''. A sequence alignment of 15 representative [[GH13]] enzymes possessing a CBM58 reveals that the location of this domain is invariant as an extension of the B domain (Figure 1). The majority of these CBM58-containing proteins are expressed within canonical polysaccharide utilization loci <cite>Foley2016 Grondin 2017</cite> of the Bacteroidetes that also encode homologs of the TonB-dependent transporter SusC and a homolog of the starch-binding protein SusD. | |
| − | + | [[File:CBM58-2.png|thumb|400px|right|'''Figure 1: CBM58 sequence alignment.''' Amino acid sequence alignment from CLUSTALW (DNASTAR, Madison, WI). All protein names (left) refer to the locus tags of CBM58-containing [[GH13]] enzymes as listed in the [http://www.cazy.org/CBM58.html CAZy database]. The numbering to the right refers to the amino acid position of each protein. For reference, the CBM58 of SusG (BT_3698) extends from residues 216-335.Red, orange and green colors at the top indicate positions of complete conservation, similarity, or more diverse sequence respectively. ]] | |
| − | |||
| − | [[File:CBM58-2.png|thumb|400px|right|'''Figure 1: CBM58 sequence alignment.''' Amino acid sequence alignment from CLUSTALW (DNASTAR, Madison, WI). All protein names (left) refer to the locus tags of CBM58-containing GH13 enzymes as listed in www.cazy.org/CBM58.html. The numbering to the right refers to the amino acid position of each protein. For reference, the CBM58 of SusG (BT_3698) extends from residues 216-335.Red, orange and green colors at the top indicate positions of complete conservation, similarity, or more diverse sequence respectively. ]] | ||
== Functionalities == | == Functionalities == | ||
| + | CBM58 is only found within [[GH13]] enzymes, and the conserved placement of the domain within the ''B. thetaiotaomicron'' SusG and like enzymes is unusual as it interrupts the fold of the catalytic domain. In the structure of SusG, the starch-binding face of CBM58 is located 45Å away and on the opposite face of the protein from the catalytic cleft <cite>Koropatkin2010</cite>. A mutant enzyme of SusG in which the CBM58 has been deleted retains catalytic activity against small substrates such as PNP-maltohexaose similar to the wild-type enzyme <cite>Koropatkin2010</cite>. The CBM-less SusG enzyme is 2-3 fold more active on autoclaved soluble starch such as maize amylopectin, but only retains ~30% of the wild-type activity on insoluble corn starch <cite>Koropatkin2010</cite>. ''B. thetaiotaomicron'' expressing only an allele for the CBM-less SusG grows the same as wild-type cells on autoclaved soluble starches <cite>Cameron2014</cite>. | ||
| − | + | The CBM58 of SusG has been replaced with the Halo-tag® protein for single-molecule fluorescence imaging of the SusG protein in live ''B. thetaiotaomicron'' cultured on glucose and starch <cite>Karunatilaka2014</cite>. In these experiments, the SusG-halo fusion partitioned between mobile and confined populations on the cell surface during growth on glucose and starch respectively, and the diffusion coefficient of the protein was measurably slower when interacting with starch granules, and in the presence of the other Sus surface proteins <cite>Karunatilaka2014</cite>. | |
| − | |||
| − | |||
== Family Firsts == | == Family Firsts == | ||
| + | ;First Identified: The CBM58 domain was identified during the biochemical and structural characterization of SusG from ''B. thetaiotaomicron'' <cite>Koropatkin2010</cite>. | ||
| − | + | ;First Structural Characterization: The crystal structure of the GH13 enzyme SusG with maltoheptaose revealed an extended B domain, CBM58, that recognizes starch and maltooligosaccharides <cite>Koropatkin2010</cite>. | |
| − | |||
| − | |||
| − | ; First Structural Characterization | ||
| − | :The crystal structure of the GH13 enzyme SusG with maltoheptaose revealed an extended B domain, CBM58, that recognizes starch and maltooligosaccharides <cite>Koropatkin2010</cite>. | ||
== References == | == References == | ||
| − | |||
<biblio> | <biblio> | ||
#Koropatkin2010 pmid=20159465 | #Koropatkin2010 pmid=20159465 | ||
| Line 54: | Line 47: | ||
#Cameron2014 pmid=25205092 | #Cameron2014 pmid=25205092 | ||
#Karunatilaka2014 pmid=25389179 | #Karunatilaka2014 pmid=25389179 | ||
| − | + | #Foley2016 pmid=27137179 | |
| + | #Grondin2017 pmid=28138099 | ||
</biblio> | </biblio> | ||
[[Category:Carbohydrate Binding Module Families|CBM058]] | [[Category:Carbohydrate Binding Module Families|CBM058]] | ||
Revision as of 16:34, 8 September 2017
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Nicole Koropatkin^^^
- Responsible Curator: ^^^Nicole Koropatkin^^^
| CAZy DB link | |
| http://www.cazy.org/CBM58.html |
Ligand specificities
Only a single CBM58 family member has been characterized, the founding member from the neopullulanase SusG of the human gut symbiont Bacteroides thetaiotaomicron [1]. The crystal structure of SusG featuring CBM58 revealed that this domain binds to maltoheptaose as well as acarbose, a pseudotetrasaccharide amylase inhibitor. Isothermal titration calorimetry with maltoheptaose and α-cyclodextrin as well as isotherm depletion assays with insoluble cornstarch further confirmed that CBM58 is a starch-specific CBM [1]. Based upon these data, the CBM58 family is believed to bind exclusively to α1,4-linked glucan structures in starch.
Structural Features
The crystal structure of CBM58 from SusG of B. thetaiotaomicron displays a canonical β-sandwich fold, with a single binding site accomodated by the loops connecting β3 and β4, β6 and β7, and β7 and β8 [1]. These loops fold over one of the β-sheets. Because CBM58 recognizes internal regions of the polypeptide chain, it can be classified as a type B CBM [2]. Like many starch-specific CBMs [3], the helical α-glucan is cradled by aromatic stacking interactions with two SusG CBM58 aromatic residues, W287 and W299, as well as hydrogen bonding interactions with K304, N330 and Y260 that aid in positioning the adjacent glucose residues that stack with the tryptophans [1].
A unique facet of the CBM58 of SusG is its position in the middle of the GH13 amylase fold, occupying residues 216-335 of the protein. This domain is inserted between α3 and β3 of the catalytic B domain (residues 153-215 and 336-363), and essentially expands the typically small B domain of GH13 enzymes. Thus the entire SusG sequence can be described from N- to C-terminus (residue numbers): A domain (43-152) - B domain (153-215) - CBM58 (215-335) - B domain (336-363) - A domain (364-607) - C domain (608-692). In SusG, two short linker sequences from residues 212-217 and 334-338, spanning 12Å between the B domain and CBM58, have B-factors and an amino acid sequence that imply this domain is held in a fixed position [1]. The CBM58 family is small, with the seminal members all arising from the Bacteroidetes phylum. This includes similar GH13 enzymes from isolates of Alistipes finegoldii, Alistipes shahii, Bacteroides dorei, Bacteroides eggerthii and Bacteroides vulgatus. A sequence alignment of 15 representative GH13 enzymes possessing a CBM58 reveals that the location of this domain is invariant as an extension of the B domain (Figure 1). The majority of these CBM58-containing proteins are expressed within canonical polysaccharide utilization loci [4, 5, 6] of the Bacteroidetes that also encode homologs of the TonB-dependent transporter SusC and a homolog of the starch-binding protein SusD.
Functionalities
CBM58 is only found within GH13 enzymes, and the conserved placement of the domain within the B. thetaiotaomicron SusG and like enzymes is unusual as it interrupts the fold of the catalytic domain. In the structure of SusG, the starch-binding face of CBM58 is located 45Å away and on the opposite face of the protein from the catalytic cleft [1]. A mutant enzyme of SusG in which the CBM58 has been deleted retains catalytic activity against small substrates such as PNP-maltohexaose similar to the wild-type enzyme [1]. The CBM-less SusG enzyme is 2-3 fold more active on autoclaved soluble starch such as maize amylopectin, but only retains ~30% of the wild-type activity on insoluble corn starch [1]. B. thetaiotaomicron expressing only an allele for the CBM-less SusG grows the same as wild-type cells on autoclaved soluble starches [7].
The CBM58 of SusG has been replaced with the Halo-tag® protein for single-molecule fluorescence imaging of the SusG protein in live B. thetaiotaomicron cultured on glucose and starch [8]. In these experiments, the SusG-halo fusion partitioned between mobile and confined populations on the cell surface during growth on glucose and starch respectively, and the diffusion coefficient of the protein was measurably slower when interacting with starch granules, and in the presence of the other Sus surface proteins [8].
Family Firsts
- First Identified
- The CBM58 domain was identified during the biochemical and structural characterization of SusG from B. thetaiotaomicron [1].
- First Structural Characterization
- The crystal structure of the GH13 enzyme SusG with maltoheptaose revealed an extended B domain, CBM58, that recognizes starch and maltooligosaccharides [1].
References
- Koropatkin NM and Smith TJ. (2010). SusG: a unique cell-membrane-associated alpha-amylase from a prominent human gut symbiont targets complex starch molecules. Structure. 2010;18(2):200-15. DOI:10.1016/j.str.2009.12.010 |
- Gilbert HJ, Knox JP, and Boraston AB. (2013). Advances in understanding the molecular basis of plant cell wall polysaccharide recognition by carbohydrate-binding modules. Curr Opin Struct Biol. 2013;23(5):669-77. DOI:10.1016/j.sbi.2013.05.005 |
- Machovic M and Janecek S. (2006). Starch-binding domains in the post-genome era. Cell Mol Life Sci. 2006;63(23):2710-24. DOI:10.1007/s00018-006-6246-9 |
- Foley MH, Cockburn DW, and Koropatkin NM. (2016). The Sus operon: a model system for starch uptake by the human gut Bacteroidetes. Cell Mol Life Sci. 2016;73(14):2603-17. DOI:10.1007/s00018-016-2242-x |
- Cameron EA, Kwiatkowski KJ, Lee BH, Hamaker BR, Koropatkin NM, and Martens EC. (2014). Multifunctional nutrient-binding proteins adapt human symbiotic bacteria for glycan competition in the gut by separately promoting enhanced sensing and catalysis. mBio. 2014;5(5):e01441-14. DOI:10.1128/mBio.01441-14 |
- Karunatilaka KS, Cameron EA, Martens EC, Koropatkin NM, and Biteen JS. (2014). Superresolution imaging captures carbohydrate utilization dynamics in human gut symbionts. mBio. 2014;5(6):e02172. DOI:10.1128/mBio.02172-14 |
- Grondin JM, Tamura K, Déjean G, Abbott DW, and Brumer H. (2017). Polysaccharide Utilization Loci: Fueling Microbial Communities. J Bacteriol. 2017;199(15). DOI:10.1128/JB.00860-16 |