CAZypedia needs your help! We have many unassigned GH, PL, CE, AA, GT, and CBM pages in need of Authors and Responsible Curators.
Scientists at all career stages, including students, are welcome to contribute to CAZypedia. Read more here, and in the 10th anniversary article in Glycobiology.
New to the CAZy classification? Read this first.
*
Consider attending the 15th Carbohydrate Bioengineering Meeting in Ghent, 5-8 May 2024.
Difference between revisions of "Glycoside Hydrolase Family 107"
Al Boraston (talk | contribs) |
Harry Brumer (talk | contribs) (small edits to mechanism section, repositioned references) |
||
| Line 29: | Line 29: | ||
== Substrate specificities == | == Substrate specificities == | ||
| − | The currently characterized [[glycoside hydrolases]] of this family are [[endo]]-acting α-fucosidases active on sulfated fucans (or fucoidans) from brown algae. All described GH107 family members are endo-1,4-fucanase of bacterial origin, and together with enzymes from the CAZY family [[GH29]], they form the clan GH-R. Sequences of GH107 family members were first reported in 2006 | + | The currently characterized [[glycoside hydrolases]] of this family are [[endo]]-acting α-fucosidases active on sulfated fucans (or fucoidans) from brown algae. All described GH107 family members are endo-1,4-fucanase of bacterial origin, and together with enzymes from the CAZY family [[GH29]], they form the clan GH-R. Sequences of GH107 family members were first reported in 2006 <cite>Colin2006</cite>, even though the enzymatic activity was reported earlier. |
== Kinetics and Mechanism == | == Kinetics and Mechanism == | ||
| − | + | The mechanism was demonstrated to be consistent with a [[classical Koshland double-displacement mechanism]] mechanism by observation of the formation of an α-''O''-linked mercaptoethanol on a L-Fuc-2,3-disulfate-(α1-3)-L-Fuc-2-sulfate disaccharide by transglycosylation <cite>Vickers2018</cite>. The same mechanism is demonstrated by [[GH29]] members, which comprise [[Clan]] GH-R together with GH107. | |
| − | The mechanism was | ||
[[Image:20191213 Gh107.png|thumb|left|800px|Figure 1: Mechanism of GH107 family. According to <cite>Vickers2018</cite>.]] | [[Image:20191213 Gh107.png|thumb|left|800px|Figure 1: Mechanism of GH107 family. According to <cite>Vickers2018</cite>.]] | ||
| Line 39: | Line 38: | ||
== Catalytic Residues == | == Catalytic Residues == | ||
| − | The catalytic nucleophile is an aspartate, while the catalytic acid-base is a histidine. The later is unusual in GHs, and a divergence from [[GH29]], but is likely necessary to avoid electronic repulsion with the substrate sulfate groups. These two residues have been identified by structural superimposition with GH29 enzymes, and are conserved within the few members of the GH107 family. The catalytic His has been further confirmed by the lack of activity of the H294Q mutant of ''Mariniflexile fucanivorans'' | + | The catalytic nucleophile is an aspartate, while the catalytic acid-base is a histidine. The later is unusual in GHs, and a divergence from [[GH29]], but is likely necessary to avoid electronic repulsion with the substrate sulfate groups. These two residues have been identified by structural superimposition with GH29 enzymes, and are conserved within the few members of the GH107 family. The catalytic His has been further confirmed by the lack of activity of the H294Q mutant of ''Mariniflexile fucanivorans'' <cite>Vickers2018</cite>. The catalytic aspartate was also proposed to be one of the catalytic residue on sequence analysis alone, in a simultaneously released paper <cite>Shultz-Johansen2018</cite>. |
== Three-dimensional structures == | == Three-dimensional structures == | ||
| − | The crystal structures of ''Mariniflexile fucanivorans'' (PDB: [{{PDBlink}}6dns 6dns],[{{PDBlink}}6dms 6dms],[{{PDBlink}}6dlh 6dlh]) and ''Psychromonas sp.'' (PDB: [{{PDBlink}}6m8n 6m8n]) have been determined in 2018 | + | The crystal structures of ''Mariniflexile fucanivorans'' (PDB: [{{PDBlink}}6dns 6dns],[{{PDBlink}}6dms 6dms],[{{PDBlink}}6dlh 6dlh]) and ''Psychromonas sp.'' (PDB: [{{PDBlink}}6m8n 6m8n]) have been determined in 2018 <cite>Vickers2018</cite>. The ''Psychromonas sp.'' (PDB: [{{PDBlink}}6m8n 6m8n]) enzyme showed a single catalytic domain with a (β/α)<sub>8</sub> / TIM-barrel fold, while in the ''Mariniflexile fucanivorans'' enzyme, this catalytic domain is followed by three Ig-like domains that wrap around the catalytic one <cite>Vickers2018</cite>. |
[[Image:GH107_full.png|500px]] | [[Image:GH107_full.png|500px]] | ||
| Line 50: | Line 49: | ||
== Family Firsts == | == Family Firsts == | ||
| − | ;First stereochemistry determination: The retaining mechanism was determined in 2018 | + | ;First stereochemistry determination: The retaining mechanism was determined in 2018 <cite>Vickers2018</cite>. |
| − | ;First catalytic nucleophile identification: An aspartic acid side chain acting as a catalytic nucleophile was identified by two simultaneous studies in the Autumn of 2018 | + | ;First catalytic nucleophile identification: An aspartic acid side chain acting as a catalytic nucleophile was identified by two simultaneous studies in the Autumn of 2018 <cite>Vickers2018 Shultz-Johansen2018</cite>. |
| − | ;First general acid/base residue identification: A histidine sidechain acting as a catalytic acid-base residue was identified in 2018 | + | ;First general acid/base residue identification: A histidine sidechain acting as a catalytic acid-base residue was identified in 2018 <cite>Vickers2018</cite>. |
| − | ;First 3-D structure: The crystal structures of ''Mariniflexile fucanivorans'' (PDB: [{{PDBlink}}6dns 6dns],[{{PDBlink}}6dms 6dms],[{{PDBlink}}6dlh 6dlh]) and ''Psychromonas sp.'' (PDB: [{{PDBlink}}6m8n 6m8n]) have been released at the same time, in 2018 | + | ;First 3-D structure: The crystal structures of ''Mariniflexile fucanivorans'' (PDB: [{{PDBlink}}6dns 6dns],[{{PDBlink}}6dms 6dms],[{{PDBlink}}6dlh 6dlh]) and ''Psychromonas sp.'' (PDB: [{{PDBlink}}6m8n 6m8n]) have been released at the same time, in 2018 <cite>Vickers2018</cite>. |
== References == | == References == | ||
Revision as of 10:03, 7 January 2020
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
- Author: ^^^David Teze^^^
- Responsible Curator: ^^^Al Boraston^^^
| Glycoside Hydrolase Family GH107 | |
| Clan | GH-R |
| Mechanism | retaining |
| Active site residues | known |
| CAZy DB link | |
| http://www.cazy.org/GH107.html | |
Substrate specificities
The currently characterized glycoside hydrolases of this family are endo-acting α-fucosidases active on sulfated fucans (or fucoidans) from brown algae. All described GH107 family members are endo-1,4-fucanase of bacterial origin, and together with enzymes from the CAZY family GH29, they form the clan GH-R. Sequences of GH107 family members were first reported in 2006 [1], even though the enzymatic activity was reported earlier.
Kinetics and Mechanism
The mechanism was demonstrated to be consistent with a classical Koshland double-displacement mechanism mechanism by observation of the formation of an α-O-linked mercaptoethanol on a L-Fuc-2,3-disulfate-(α1-3)-L-Fuc-2-sulfate disaccharide by transglycosylation [2]. The same mechanism is demonstrated by GH29 members, which comprise Clan GH-R together with GH107.
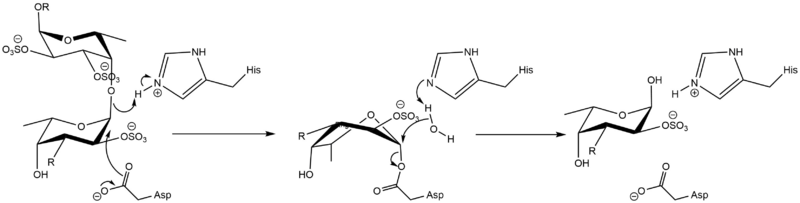
Catalytic Residues
The catalytic nucleophile is an aspartate, while the catalytic acid-base is a histidine. The later is unusual in GHs, and a divergence from GH29, but is likely necessary to avoid electronic repulsion with the substrate sulfate groups. These two residues have been identified by structural superimposition with GH29 enzymes, and are conserved within the few members of the GH107 family. The catalytic His has been further confirmed by the lack of activity of the H294Q mutant of Mariniflexile fucanivorans [2]. The catalytic aspartate was also proposed to be one of the catalytic residue on sequence analysis alone, in a simultaneously released paper [3].
Three-dimensional structures
The crystal structures of Mariniflexile fucanivorans (PDB: 6dns,6dms,6dlh) and Psychromonas sp. (PDB: 6m8n) have been determined in 2018 [2]. The Psychromonas sp. (PDB: 6m8n) enzyme showed a single catalytic domain with a (β/α)8 / TIM-barrel fold, while in the Mariniflexile fucanivorans enzyme, this catalytic domain is followed by three Ig-like domains that wrap around the catalytic one [2].
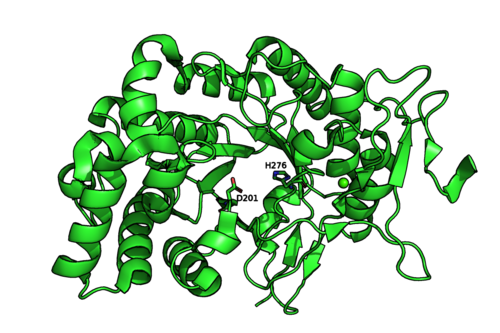
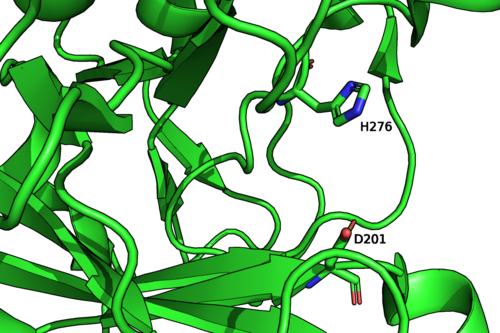
Figure 2: Psychromonas sp. GH107 structure and catalytic residues (PDB: 6m8n).
Family Firsts
- First stereochemistry determination
- The retaining mechanism was determined in 2018 [2].
- First catalytic nucleophile identification
- An aspartic acid side chain acting as a catalytic nucleophile was identified by two simultaneous studies in the Autumn of 2018 [2, 3].
- First general acid/base residue identification
- A histidine sidechain acting as a catalytic acid-base residue was identified in 2018 [2].
- First 3-D structure
- The crystal structures of Mariniflexile fucanivorans (PDB: 6dns,6dms,6dlh) and Psychromonas sp. (PDB: 6m8n) have been released at the same time, in 2018 [2].
References
- Colin S, Deniaud E, Jam M, Descamps V, Chevolot Y, Kervarec N, Yvin JC, Barbeyron T, Michel G, and Kloareg B. (2006). Cloning and biochemical characterization of the fucanase FcnA: definition of a novel glycoside hydrolase family specific for sulfated fucans. Glycobiology. 2006;16(11):1021-32. DOI:10.1093/glycob/cwl029 |
- Vickers C, Liu F, Abe K, Salama-Alber O, Jenkins M, Springate CMK, Burke JE, Withers SG, and Boraston AB. (2018). Endo-fucoidan hydrolases from glycoside hydrolase family 107 (GH107) display structural and mechanistic similarities to α-l-fucosidases from GH29. J Biol Chem. 2018;293(47):18296-18308. DOI:10.1074/jbc.RA118.005134 |
- Schultz-Johansen M, Cueff M, Hardouin K, Jam M, Larocque R, Glaring MA, Hervé C, Czjzek M, and Stougaard P. (2018). Discovery and screening of novel metagenome-derived GH107 enzymes targeting sulfated fucans from brown algae. FEBS J. 2018;285(22):4281-4295. DOI:10.1111/febs.14662 |