CAZypedia needs your help! We have many unassigned GH, PL, CE, AA, GT, and CBM pages in need of Authors and Responsible Curators.
Scientists at all career stages, including students, are welcome to contribute to CAZypedia. Read more here, and in the 10th anniversary article in Glycobiology.
New to the CAZy classification? Read this first.
*
Consider attending the 15th Carbohydrate Bioengineering Meeting in Ghent, 5-8 May 2024.
Glycoside Hydrolase Family 65
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
- Author: ^^^Hiroyuki Nakai^^^
- Responsible Curator: ^^^Hiroyuki Nakai^^^
| Glycoside Hydrolase Family GH65 | |
| Clan | GH-L |
| Mechanism | inverting |
| Active site residues | known |
| CAZy DB link | |
| http://www.cazy.org/GH65.html | |
Substrate specificities
Glycoside hydrolases belonging to GH65 act on substrates containing α-glucosidic linkages. GH65 contains mainly phosphorylases; maltose (Glc-α-1,4-Glc) phosphorylase (EC 2.4.1.8), trehalose (Glc-α1,α1-Glc) phosphorylase (EC 2.4.1.64), kojibiose (Glc-α-1,2-Glc) phosphorylase (EC 2.4.1.230), and trehalose 6-phosphate (Glc-α1,α1-Glc6P) phosphorylase (EC 2.4.1.-). Notably α,α-trehalases (EC 3.2.1.28), which a glycoside hydrolases, are also GH65 members.
Kinetics and Mechanism
Phosphorolysis by GH65 enzymes proceeds with inversion of anomeric configuration, as first shown by Fitting and Doudoroff [1] using maltose phosphorylase from Neisseria meningitidis, i.e. the reaction catalyzed is: maltose + Pi ↔ β-glucose 1-phosphate + glucose. The reaction mechanism for inverting GH65 phosphorylase has been proposed to be similar to a general acid/base-catalysed one-step displacement mechanism for inverting glycoside hydrolases [2, 3]. This mechanism involves direct nucleophilic attack by phosphate on the anomeric (C1) carbon assisted by general acid catalysis involving protonation of the glycosidic oxygen. In this mechanism, phosphate is the nucleophile, rather than a water molecule activated by a general base catalyst as in inverting glycoside hydrolases. The possibility of general base assistance in the phosphorylases of this family, via a histidine residue proximal to the phosphate nucleophile, is also suggested [2]. The inverting phosphorolysis catalyzed by GH65 enzyme is readily reversible, which confers the phosphorylases with a capacity to effectively synthesize various α-glucosides from β-glucose 1-phosphate as a glycosyl donor, together with various acceptor molecules. Notably β-glucosyl fluoride can also be used as donor in synthetic reactions [4].
Catalytic Residues
The general acid catalyst was first predicted by superimposing the active site structure of maltose phosphorylase from Lactobacillus brevis [2] with the catalytic (α/α)6 barrel domain of a GH15 glucoamylase (EC 3.2.1.28) from Aspergillus awamori [5]. Considering the similarities of the active site structure, Glu487 of L. brevis maltose phosphorylase was proposed to be the general acid. Additional support for this assignment was obtained by site-direct mutagenesis of Glu487 of Paenibacillus sp. maltose phosphorylase [6], which corresponds to the Glu487 of L. brevis maltose phosphorylase. A histidine located near the phosphate (His671 in L. brevis maltose phosphorylase) may assist catalysis by acting as a general base [2], but no experimental evidence has yet been provided.
Three-dimensional structures
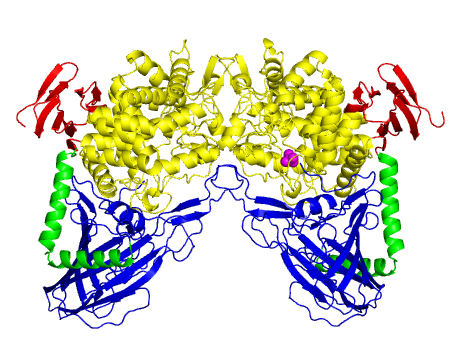
The three-dimensional structure of L. brevis maltose phosphorylase (PDB ID 1h54) represents the first determined in this family [2]. L. brevis maltose phosphorylase is a dimeric enzyme (Figure 1). The monomer structure consists of an N-terminal complex β sandwich domain, a helical linker, an (α/α)6 barrel catalytic domain, and a C-terminal β sheet domain. The structure of the (α/α)6 barrel domain is similar to those of GH15 glucoamylases (EC 3.2.1.28) [5]. GH15 and GH65 together constitute glycoside hydrolase Clan GH-L [7]. GH94 phosphorylases and a GH95 fucosidase were also shown to be structurally similar to GH65 [8, 9].
Family Firsts
- First stereochemistry determination
- Maltose phosphorylase (EC 2.4.1.8) from Neisseria meningitidis [1].
- First sequence identification
- Maltose phosphorylase (EC 2.4.1.8) from Lactobacillus sanfranciscensis DSM 20451T [10]
- Trehalose phosphorylase (EC 2.4.1.64) from Thermoanaerobacter brockii ATCC 35047 [11]
- Kojibiose phosphorylase (EC 2.4.1.230) from Thermoanaerobacter brockii ATCC 35047 [12]
- Trehalose 6-phosphate phosphorylase (EC 2.4.1.-) from Lactococcus lactis ssp. lactis 19435 [13]
- α,α-Trehalase (EC 3.2.1.28) from Saccharomyces cerevisiae S288C [14].
- First general acid residue identification
- Maltose phosphorylase (EC 2.4.1.8) from Lactobacillus brevis by X-ray structure analysis [2] and confirmed by mutagenesis for Paenibacillus sp. maltose phosphorylase (EC 2.4.1.8) [6].
- First three-dimentional structure determination
- Maltose phosphorylase (EC 2.4.1.8) from Lactobacillus brevis [2].
References
- FITTING C and DOUDOROFF M. (1952). Phosphorolysis of maltose by enzyme preparations from Neisseria meningitidis. J Biol Chem. 1952;199(1):153-63. | Google Books | Open Library
- Egloff MP, Uppenberg J, Haalck L, and van Tilbeurgh H. (2001). Crystal structure of maltose phosphorylase from Lactobacillus brevis: unexpected evolutionary relationship with glucoamylases. Structure. 2001;9(8):689-97. DOI:10.1016/s0969-2126(01)00626-8 |
- Nakai H, Baumann MJ, Petersen BO, Westphal Y, Schols H, Dilokpimol A, Hachem MA, Lahtinen SJ, Duus JØ, and Svensson B. (2009). The maltodextrin transport system and metabolism in Lactobacillus acidophilus NCFM and production of novel alpha-glucosides through reverse phosphorolysis by maltose phosphorylase. FEBS J. 2009;276(24):7353-65. DOI:10.1111/j.1742-4658.2009.07445.x |
- Tsumuraya Y, Brewer CF, and Hehre EJ. (1990). Substrate-induced activation of maltose phosphorylase: interaction with the anomeric hydroxyl group of alpha-maltose and alpha-D-glucose controls the enzyme's glucosyltransferase activity. Arch Biochem Biophys. 1990;281(1):58-65. DOI:10.1016/0003-9861(90)90412-r |
- Aleshin A, Golubev A, Firsov LM, and Honzatko RB. (1992). Crystal structure of glucoamylase from Aspergillus awamori var. X100 to 2.2-A resolution. J Biol Chem. 1992;267(27):19291-8. DOI:10.2210/pdb1gly/pdb |
-
Hidaka Y, Hatada Y, Akita M, Yoshida M, Nakamura N, Takada M, Nakakuki T, Ito S, and Horikoshi K. Maltose phosphorylase from a deep-sea Paenibacillus sp.: Enzymatic properties and nucleotide and amino-acid sequences. Enzyme and Microbial Technology, Volume 37, Issue 2, 1 July 2005, Pages 185-194. doi:10.1016/j.enzmictec.2005.02.010
- Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, and Henrissat B. (2009). The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res. 2009;37(Database issue):D233-8. DOI:10.1093/nar/gkn663 |
- Hidaka M, Honda Y, Kitaoka M, Nirasawa S, Hayashi K, Wakagi T, Shoun H, and Fushinobu S. (2004). Chitobiose phosphorylase from Vibrio proteolyticus, a member of glycosyl transferase family 36, has a clan GH-L-like (alpha/alpha)(6) barrel fold. Structure. 2004;12(6):937-47. DOI:10.1016/j.str.2004.03.027 |
- Nagae M, Tsuchiya A, Katayama T, Yamamoto K, Wakatsuki S, and Kato R. (2007). Structural basis of the catalytic reaction mechanism of novel 1,2-alpha-L-fucosidase from Bifidobacterium bifidum. J Biol Chem. 2007;282(25):18497-18509. DOI:10.1074/jbc.M702246200 |
- Ehrmann MA and Vogel RF. (1998). Maltose metabolism of Lactobacillus sanfranciscensis: cloning and heterologous expression of the key enzymes, maltose phosphorylase and phosphoglucomutase. FEMS Microbiol Lett. 1998;169(1):81-6. DOI:10.1111/j.1574-6968.1998.tb13302.x |
- Maruta K, Mukai K, Yamashita H, Kubota M, Chaen H, Fukuda S, and Kurimoto M. (2002). Gene encoding a trehalose phosphorylase from Thermoanaerobacter brockii ATCC 35047. Biosci Biotechnol Biochem. 2002;66(9):1976-80. DOI:10.1271/bbb.66.1976 |
- Yamamoto T, Maruta K, Mukai K, Yamashita H, Nishimoto T, Kubota M, Fukuda S, Kurimoto M, and Tsujisaka Y. (2004). Cloning and sequencing of kojibiose phosphorylase gene from Thermoanaerobacter brockii ATCC35047. J Biosci Bioeng. 2004;98(2):99-106. DOI:10.1016/S1389-1723(04)70249-2 |
- Andersson U, Levander F, and Rådström P. (2001). Trehalose-6-phosphate phosphorylase is part of a novel metabolic pathway for trehalose utilization in Lactococcus lactis. J Biol Chem. 2001;276(46):42707-13. DOI:10.1074/jbc.M108279200 |
- Destruelle M, Holzer H, and Klionsky DJ. (1995). Isolation and characterization of a novel yeast gene, ATH1, that is required for vacuolar acid trehalase activity. Yeast. 1995;11(11):1015-25. DOI:10.1002/yea.320111103 |