CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 47
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH47 | |
| Clan | none, (α/α)7 fold |
| Mechanism | inverting |
| Active site residues | debated |
| CAZy DB link | |
| https://www.cazy.org/GH47.html | |
Substrate specificities
GH47 glycoside hydrolases are exo-acting α-1,2-mannosidases. Members from this family play important roles in the processing of N-glycans and are classified as Class I mannosidases; Class II mannosidases refer to those of family GH38 [1]. Three subfamilies of GH47 enzymes have been identified based upon their different substrate specifities.
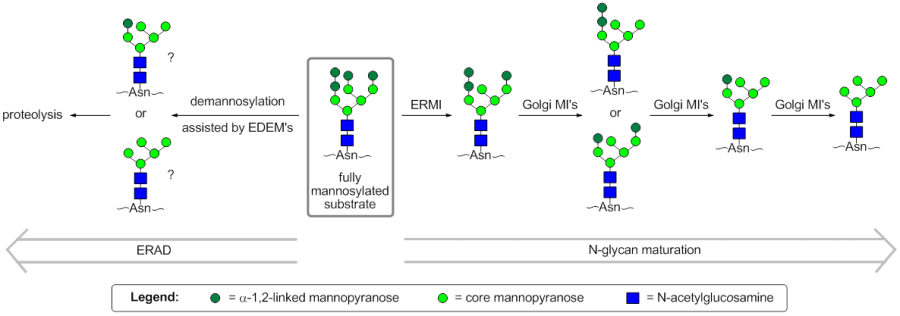
In mammals, ER-α-mannosidase I (ERMI) is representative of the GH47 subfamily that acts upon Man9GlcNAc2 to cleave a mannose from the B-chain to afford Man8GlcNAc2. Extended incubation results in further demannosylated products in vitro [2], as does overexpression in vivo [3]. Pulse-chase studies have found that Saccharomyces cerevisiae α-mannosidase I, the only GH47 mannosidase of the organism, bears essentially the same activity as mammalian ER-α-mannosidase I [1].
In mammals, the GH47 Golgi mannosidase I (Golgi MI) subfamily acts on Man8-9GlcNAc2 to afford Man5GlcNAc2 and is composed of 3 members (denoted IA, IB and IC) [4]. In contrast to mammalian ER-α-mannosidase I, the Golgi-resident GH47 mannosidases preferentially cleave from the A- and C-chains of the glycan in an order that depends on the subfamily member [5]. All mammalian Golgi mannosidase I enzymes tested thus far have relatively low activity against the B-chain of the glycan, meaning that GH47 mannosidases from the ER and Golgi have complementary actitivities.
The third GH47 subfamily is composed of the ER degradation-enhancing mannosidase-like (EDEM) proteins. This subfamily contains 3 members in humans and was initially believed to not have direct glycosidase activity. However, it now appears that the EDEM1 and EDEM3 isoforms have glycosidase activity in vivo [6, 7]. It has been suggested that the EDEM proteins act as cofactors, increasing the activity of ERMI [8]. All of the EDEM isoforms accelerate the disposal of terminally misfolded proteins through ER-associated degradation (ERAD) [7, 9, 10]. However, the process of recognition of terminally misfolded proteins and the role of EDEM proteins in ERAD is not fully understood. A current model for the early stages of ERAD states that correct folding mediated by the calnexin folding cycle must occur before the slow demannosylation of the substrate affords Man6GlcNAc2, which is no longer a substrate for reglucosylation by UGGT1 and re-entry into the calnexin folding cycle [8]. It is not clear whether this extensive demannosylation is performed solely by ERMI in vivo, which is found in high concentrations in the ER-derived quality control compartment, or if it is also performed by Golgi MI's and EDEM's.
Fewer studies have focussed upon the role of GH47 enzymes in plants. However, it has been found that these mannosidases are essential for N-glycan processing in Arabidopsis thaliana [11].
A bacterial GH47 enzyme from Caulobacter strain K31 was active on a range of aryl α-D-mannosides; its activity on N-glycans was not reported [12].
Kinetics and Mechanism
GH47 mannosidases catalyze glycosidic cleavage with inversion of stereochemistry, as first determined employing 1H NMR spectroscopy with Saccharomyces cervisiae α-1,2-mannosidase using Man9GlcNAc as a substrate [13]. Classical inverting glycosidases operate through a single displacement mechanism, where a general base residue acts to deprotonate a water molecule, facilitating nucleophilic attack at the anomeric position. This is assisted by concurrent activation of the glycosidic linkage through protonation by a general acid residue.
GH47 enzymes are Ca2+-dependent, as demonstrated by loss of activity upon addition of the metal binding ligand EDTA, and restoration of activity through subsequent addition of Ca2+ [14]. Exo-α-mannosidases from GH38 and GH92 also require a metal ion for catalysis.
GH47 mannosidases operate through an unusual 3,OB/3S1→3H4‡→1C4 conformational itinerary. Structural studies employing unhydrolysable S-linked substrate analogues have examined the Michaelis complex, with the ligands found to bind in 3S1 [15] and 3,OB/3S1 conformations [12]. Mannoimidazole, whose binding to other mannosidases has been shown to be consistent with good transition state mimicry [16], binds GH47 in a 3H4 conformation [12]. Noeuromycin [12], kifunensine [17] and 1-deoxymannojirimycin [17] all bind in a 1C4 conformation, analogous to enzyme-product complexes. Computational studies also support a 3,OB/3S1→3H4‡→1C4 conformational itinerary [12, 18, 19]. Quantum mechanical/molecular modelling calculations have found that the free energy landscape of α-D-mannopyranose is perturbed on-enzyme such that the accessible conformations of the ligand are altered to those that correlate well with a 3,OB/3S1→3H4‡→1C4 conformational itinerary [12].
Catalytic Residues
Unequivocal assignment of catalytic residues for GH47 α-mannosidases is complicated by the presence of 3 carboxylate-containing residues all approximately 9.5 Å apart from one another in the active site. Each of these could plausibly fulfill roles as catalytic residues [20]. Furthermore, all of the plausible catalytic residues complex water, as would be expected of the general base residue. Thus, it appears that the general acid residue transmits a proton to the glycosidic oxygen atom through a water molecule. Site-directed mutagenesis of residues in the α-mannosidase I of Aspergillus saitoi and Saccharomyces cerevisiae predated determination of a crystal structure but demonstrated that mutation of any of the three catalytic candidates led to total or near-total loss of activity [21, 22]. Mutagenesis of residues in human ER α-mannosidase I, informed by the determination of the crystal structure, could not unambiguously assign the role of catalytic residues [15]. Glu132 (Glu330 in human ER α-mannosidase I) in Saccharomyces cerevisiae α-mannosidase I was initially thought to be most likely candidate as the general base residue [20]. Subsequent crystal structures of human ER α-mannosidase I in complex with kifunensine and 1-deoxymannojirimycin bound these ligands in an unusual 1C4 conformation [17]. These complexes were interpreted as being representative of a 1C4 Michaelis complex, making Glu330 (Glu132 in Saccharomyces) incompatible with a role acting as the general base in an inverting mechanism. Thus, the general base residue was reassigned as either Glu599 or Asp463 (Glu435 and Asp275 in Saccharomyces, respectively). A computational docking study found Glu599 to be the most likely general base, with Ca2+ also coordinated to the nucleophilic water molecule [23]. However, complexes with S-linked substrate analogues implicate a 3,OB/3S1→3H4‡→1C4 conformational itinerary, the reverse of that used to preclude Glu330 (Glu132 in Saccharomyces) as the general base residue [12, 15]. The position of Glu330 (Glu132 in Saccharomyces) on the opposite face of the glycan ring to the putative general base residue, Glu599 in human ER α-mannosidase I (Glu435 in Saccharomyces), is consistent with a role as the general acid [17]. Arg334 is within ion-pairing distance to Glu330 and coordinates to the same water molecule, suggestive of a possible catalytic zwitterionic arginine-carboxylate dyad [15]. However, a computational docking study found Asp463 (Asp275 in Saccharomyces) to be the most likely general acid, based upon the assumption that GH47 mannosidases are anti-protonators [24]. The low nanomolar binding of mannoimidazole to CkGH47 is consistent with anti-protonation [12].
Three-dimensional structures
GH47 enzymes adopt a (α/α)7 barrel fold with a Ca2+ ion coordinated at the base of the barrel that is plugged by a β-hairpin at the C-terminus [20]. The –1 subsite lies in the core of the barrel with Ca2+ coordinating to the 2-OH and 3-OH groups of a ligand (inhibitor or substrate analogue), whose glycan ring is parallel to the barrel upon complexation [17].
The structural basis for differences in N-glycan branch specificity between ER and Golgi GH47 α-mannosidases has been examined through crystallographic studies comparing their binding to N-glycans [25]. The presumed enzyme-product complexes differed in their oligosaccharide conformation such that different oligosaccharide branches, corresponding to those readily cleaved by the respective enzymes, were projected into the active site.
| Three-dimensional structure of human GH47 α-mannosidase, PDB code [1] [4]. | Three-dimensional structure of human GH47 α-mannosidase in complex with 1-deoxymannojirimycin, PDB code [2] [4]. |
|---|---|
|
<jmol> <jmolApplet> <color>white</color> <frame>true</frame> <uploadedFileContents>1FMI.pdb</uploadedFileContents> <script>cpk off; wireframe off; cartoon; color cartoon powderblue; select ligand; wireframe 0.3; select MG; spacefill; set spin Y 10; spin off; set antialiasDisplay OFF</script> </jmolApplet> </jmol> |
<jmol> <jmolApplet> <color>white</color> <frame>true</frame> <uploadedFileContents>1FO2.pdb</uploadedFileContents> <script>cpk off; wireframe off; cartoon; color cartoon powderblue; select DMJ; wireframe 0.3; set spin Y 10; spin off; set antialiasDisplay OFF</script> </jmolApplet> </jmol> |
Family Firsts
- First sterochemistry determination
- Saccharomyces cerevisiae α-1,2-mannosidase was shown to be inverting by 1H NMR [13].
- First general base identification
- Unambiguous identification hindered by presence of 3 carboxylate-containing residues in the active site that coordinate ligands through water molecules [20]. Believed to be Glu559 in human ER α-mannosidase I (Glu435 in S. cerevisiae) [23].
- First general acid identification
- Unambiguous identification hindered by presence of 3 carboxylate-containing residues in the active site that coordinate ligands through water molecules [20]. Reported to be Glu330 in human ER α-mannosidase I (Glu132 in S. cerevisiae) [17], however, a computational study has concluded that Asp463 acts as the general acid in human ER α-mannosidase I (Asp275 in S. cerevisiae) [24].
- First 3-D structure
- Saccharomyces cerevisiae α-1,2-mannosidase [20].
References
- Herscovics A (2001). Structure and function of Class I alpha 1,2-mannosidases involved in glycoprotein synthesis and endoplasmic reticulum quality control. Biochimie. 2001;83(8):757-62. DOI:10.1016/s0300-9084(01)01319-0 |
- Herscovics A, Romero PA, and Tremblay LO. (2002). The specificity of the yeast and human class I ER alpha 1,2-mannosidases involved in ER quality control is not as strict previously reported. Glycobiology. 2002;12(4):14G-15G. | Google Books | Open Library
- Hosokawa N, Tremblay LO, You Z, Herscovics A, Wada I, and Nagata K. (2003). Enhancement of endoplasmic reticulum (ER) degradation of misfolded Null Hong Kong alpha1-antitrypsin by human ER mannosidase I. J Biol Chem. 2003;278(28):26287-94. DOI:10.1074/jbc.M303395200 |
- Tremblay LO and Herscovics A. (2000). Characterization of a cDNA encoding a novel human Golgi alpha 1, 2-mannosidase (IC) involved in N-glycan biosynthesis. J Biol Chem. 2000;275(41):31655-60. DOI:10.1074/jbc.M004935200 |
- Lal A, Pang P, Kalelkar S, Romero PA, Herscovics A, and Moremen KW. (1998). Substrate specificities of recombinant murine Golgi alpha1, 2-mannosidases IA and IB and comparison with endoplasmic reticulum and Golgi processing alpha1,2-mannosidases. Glycobiology. 1998;8(10):981-95. DOI:10.1093/glycob/8.10.981 |
- Hosokawa N, Tremblay LO, Sleno B, Kamiya Y, Wada I, Nagata K, Kato K, and Herscovics A. (2010). EDEM1 accelerates the trimming of alpha1,2-linked mannose on the C branch of N-glycans. Glycobiology. 2010;20(5):567-75. DOI:10.1093/glycob/cwq001 |
- Hirao K, Natsuka Y, Tamura T, Wada I, Morito D, Natsuka S, Romero P, Sleno B, Tremblay LO, Herscovics A, Nagata K, and Hosokawa N. (2006). EDEM3, a soluble EDEM homolog, enhances glycoprotein endoplasmic reticulum-associated degradation and mannose trimming. J Biol Chem. 2006;281(14):9650-8. DOI:10.1074/jbc.M512191200 |
- Lederkremer GZ (2009). Glycoprotein folding, quality control and ER-associated degradation. Curr Opin Struct Biol. 2009;19(5):515-23. DOI:10.1016/j.sbi.2009.06.004 |
- Hosokawa N, Wada I, Hasegawa K, Yorihuzi T, Tremblay LO, Herscovics A, and Nagata K. (2001). A novel ER alpha-mannosidase-like protein accelerates ER-associated degradation. EMBO Rep. 2001;2(5):415-22. DOI:10.1093/embo-reports/kve084 |
- Olivari S, Galli C, Alanen H, Ruddock L, and Molinari M. (2005). A novel stress-induced EDEM variant regulating endoplasmic reticulum-associated glycoprotein degradation. J Biol Chem. 2005;280(4):2424-8. DOI:10.1074/jbc.C400534200 |
- Liebminger E, Hüttner S, Vavra U, Fischl R, Schoberer J, Grass J, Blaukopf C, Seifert GJ, Altmann F, Mach L, and Strasser R. (2009). Class I alpha-mannosidases are required for N-glycan processing and root development in Arabidopsis thaliana. Plant Cell. 2009;21(12):3850-67. DOI:10.1105/tpc.109.072363 |
- Thompson AJ, Dabin J, Iglesias-Fernández J, Ardèvol A, Dinev Z, Williams SJ, Bande O, Siriwardena A, Moreland C, Hu TC, Smith DK, Gilbert HJ, Rovira C, and Davies GJ. (2012). The reaction coordinate of a bacterial GH47 α-mannosidase: a combined quantum mechanical and structural approach. Angew Chem Int Ed Engl. 2012;51(44):10997-1001. DOI:10.1002/anie.201205338 |
- Lipari F, Gour-Salin BJ, and Herscovics A. (1995). The Saccharomyces cerevisiae processing alpha 1,2-mannosidase is an inverting glycosidase. Biochem Biophys Res Commun. 1995;209(1):322-6. DOI:10.1006/bbrc.1995.1506 |
- Jelinek-Kelly S and Herscovics A. (1988). Glycoprotein biosynthesis in Saccharomyces cerevisiae. Purification of the alpha-mannosidase which removes one specific mannose residue from Man9GlcNAc. J Biol Chem. 1988;263(29):14757-63. | Google Books | Open Library
- Karaveg K, Siriwardena A, Tempel W, Liu ZJ, Glushka J, Wang BC, and Moremen KW. (2005). Mechanism of class 1 (glycosylhydrolase family 47) {alpha}-mannosidases involved in N-glycan processing and endoplasmic reticulum quality control. J Biol Chem. 2005;280(16):16197-207. DOI:10.1074/jbc.M500119200 |
- Tailford LE, Offen WA, Smith NL, Dumon C, Morland C, Gratien J, Heck MP, Stick RV, Blériot Y, Vasella A, Gilbert HJ, and Davies GJ. (2008). Structural and biochemical evidence for a boat-like transition state in beta-mannosidases. Nat Chem Biol. 2008;4(5):306-12. DOI:10.1038/nchembio.81 |
- Vallee F, Karaveg K, Herscovics A, Moremen KW, and Howell PL. (2000). Structural basis for catalysis and inhibition of N-glycan processing class I alpha 1,2-mannosidases. J Biol Chem. 2000;275(52):41287-98. DOI:10.1074/jbc.M006927200 |
- Mulakala C, Nerinckx W, and Reilly PJ. (2006). Docking studies on glycoside hydrolase Family 47 endoplasmic reticulum alpha-(1-->2)-mannosidase I to elucidate the pathway to the substrate transition state. Carbohydr Res. 2006;341(13):2233-45. DOI:10.1016/j.carres.2006.05.011 |
- Mulakala C, Nerinckx W, and Reilly PJ. (2007). The fate of beta-D-mannopyranose after its formation by endoplasmic reticulum alpha-(1-->2)-mannosidase I catalysis. Carbohydr Res. 2007;342(2):163-9. DOI:10.1016/j.carres.2006.11.012 |
- Vallée F, Lipari F, Yip P, Sleno B, Herscovics A, and Howell PL. (2000). Crystal structure of a class I alpha1,2-mannosidase involved in N-glycan processing and endoplasmic reticulum quality control. EMBO J. 2000;19(4):581-8. DOI:10.1093/emboj/19.4.581 |
- Lipari F and Herscovics A. (1999). Calcium binding to the class I alpha-1,2-mannosidase from Saccharomyces cerevisiae occurs outside the EF hand motif. Biochemistry. 1999;38(3):1111-8. DOI:10.1021/bi981643i |
- Fujita A, Yoshida T, and Ichishima E. (1997). Five crucial carboxyl residues of 1,2-alpha-mannosidase from Aspergillus saitoi (A. phoenicis), a food microorganism, are identified by site-directed mutagenesis. Biochem Biophys Res Commun. 1997;238(3):779-83. DOI:10.1006/bbrc.1997.7389 |
- Mulakala C and Reilly PJ. (2002). Understanding protein structure-function relationships in Family 47 alpha-1,2-mannosidases through computational docking of ligands. Proteins. 2002;49(1):125-34. DOI:10.1002/prot.10206 |
- Cantú D, Nerinckx W, and Reilly PJ. (2008). Theory and computation show that Asp463 is the catalytic proton donor in human endoplasmic reticulum alpha-(1-->2)-mannosidase I. Carbohydr Res. 2008;343(13):2235-42. DOI:10.1016/j.carres.2008.05.026 |
- Tempel W, Karaveg K, Liu ZJ, Rose J, Wang BC, and Moremen KW. (2004). Structure of mouse Golgi alpha-mannosidase IA reveals the molecular basis for substrate specificity among class 1 (family 47 glycosylhydrolase) alpha1,2-mannosidases. J Biol Chem. 2004;279(28):29774-86. DOI:10.1074/jbc.M403065200 |