CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Difference between revisions of "Glycoside Hydrolase Family 194"
| (10 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
<!-- RESPONSIBLE CURATORS: Please replace the {{UnderConstruction}} tag below with {{CuratorApproved}} when the page is ready for wider public consumption --> | <!-- RESPONSIBLE CURATORS: Please replace the {{UnderConstruction}} tag below with {{CuratorApproved}} when the page is ready for wider public consumption --> | ||
| − | {{ | + | {{CuratorApproved}} |
* [[Author]]: [[User:Masahiro Nakajima|Masahiro Nakajima]] | * [[Author]]: [[User:Masahiro Nakajima|Masahiro Nakajima]] | ||
* [[Responsible Curator]]: [[User:Masahiro Nakajima|Masahiro Nakajima]] | * [[Responsible Curator]]: [[User:Masahiro Nakajima|Masahiro Nakajima]] | ||
| Line 29: | Line 29: | ||
== Substrate specificities == | == Substrate specificities == | ||
| − | PgSGL3(H744_1c0222, KEGG) from <i>Photobacterium gaetbulicola</i> | + | PgSGL3(H744_1c0222, KEGG) from <i>Photobacterium gaetbulicola</i> was characterized as reported in 2025 <cite>Nakajima2025</cite>. The enzyme specifically hydrolyzes β-1,2-glucan to produce β-1,2-glucooligosaccharides in an endolytic manner. PgSGL3 preferentially produces β-1,2-glucooctasaccharide at the initial stage of hydrolysis of β-1,2-glucan. |
== Kinetics and Mechanism == | == Kinetics and Mechanism == | ||
| − | PgSGL3 follows anomer-inverting mechanism | + | Hydrolysis of β-1,2-glucan by PgSGL3 suggests that the enzyme follows anomer-[[inverting]] mechanism <cite>Nakajima2025</cite>. Analysis of the change of the degree of optical rotation during hydrolysis of β-1,2-glucan and after addition of aqueous ammonia. Sharp decrease of the degree of optical rotation by aqueous ammonia is the same pattern as in the case of [[GH162]] β-1,2-glucanase from <i>Talaromyces funiculosus</i> (TfSGL), an anomer-[[inverting]] enzyme <cite>Tanaka2019</cite>. |
== Catalytic Residues == | == Catalytic Residues == | ||
| − | E214(PgSGL3) is the putative general acid as this residue is structurally well-superimposed with the general acid (E262) in [[GH162]] | + | E214(PgSGL3) is the putative general acid as this residue is structurally well-superimposed with the general acid (E262) in [[GH162]] TfSGL [2]. E214Q mutant shows drastic decrease in catalytic activity compared to the wild-type enzyme [1]. E214 is also conserved across other GH-S clan families including [[GH144]], [[GH192]], and [[GH193]]. In [[GH189]], a family related to clan GH-S, this equivalent residue acts as a catalytic acid/base <cite>Tanaka2024</cite><br> |
| + | Similarly, D148 (PgSGL3) is a residue conserved spatially with several β-1,2-glucanases; [[GH144]] (from <i>Chitinophaga pinensis</i> and <i>Xanthomonas campestris</i> pv. <i>campestris</i>), [[GH192]] (from <i>P. gaetbulicala</i>), and [[GH193]] (from <i>Sanguibacter keddieii</i>) <cite>Nakajima2025, Abe2017</cite>. D148N mutant also shows drastically decreased activity against the wild-type enzyme. Mutational analysis alone is insufficient to definitively identify catalytic residues because a reaction mechanism of [[GH194]] is atypical. | ||
| + | |||
| + | A plausible substrate binding mode of PgSGL3 can be obtained by superimposed with the complex structure of [[GH144]] β-1,2-glucanase from <i>X. campestris</i> pv. <i>campestris</i> with β-1,2-glucoheptaose. However, no nucleophilic water is observed and no clear pathway for proton transfer from a nucleophilic water to a general base can be traced. It should be noted that the position of D148 (PgSGL3) does not correspond to that of the general base in [[GH162]] TfSGL nor to the nucleophile in [[GH189]] β-1,2-glucanotransferase <cite>Nakajima2025, Tanaka2019, Tanaka2024</cite>, which suggests a difference in reaction mechanism between these families. | ||
== Three-dimensional structures == | == Three-dimensional structures == | ||
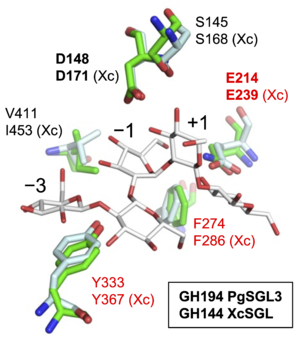
| − | A ligand-free structure of PgSGL3 is available <cite> | + | [[Image:GH194_pedia.png|thumb|300px|'''Fig. 1. Superimposition of PgSGL3([[GH194]]) and XcSGL([[GH144]])''' PgSGL3(PDB ID, [{{PDBlink}}8XUK 8XUK]) and XcSGL(PDB ID, [{{PDBlink}}8XUL 8XUL]) are shown in light cyan and green, respectively. Residues labelled in red are the SGL-defining residues. Residues of XcSGL are labelled with Xc. This figure is modified from <cite>Nakajima2025</cite>]]A ligand-free structure of PgSGL3 is available (PDB ID, [{{PDBlink}} 8XUK]) <cite>Nakajima2025</cite>. PgSGL3 is composed of a single (α/α)<sub>6</sub>-barrel fold. The overall structure and the shape of catalytic pocket of PgSGL3 are similar to those of [[GH144]] β-1,2-glucanases. The two candidate catalytic residues described above are well-superimposed with [[GH144]] β-1,2-glucanases. Based on the similarity, [[GH194]] is classified into clan GH-S, the same clan as [[GH144]]. Clan GH-S is composed of [[GH144]], [[GH162]], [[GH192]], [[GH193]], and [[GH194]] which are distantly related families. Although [[GH189]] is also a family distantly related to these families, [[GH189]] is excluded from clan GH-S due to the difference in the reaction mechanism ([[GH189]] enzymes follow anomer-[[retaining]] mechanism). These distantly related families including [[GH189]] is called '''SGL clan''' <cite>Nakajima2025</cite>. The three residues labelled in red in the catalytic pocket (Fig. 1) are conserved within [[GH144]], [[GH192]] and [[GH193]] families. The three conserved residues are considered as residues defining the SGL clan. Among the three residues, the glutamate residue is the candidate general acid described above. |
== Family Firsts == | == Family Firsts == | ||
| − | ;First stereochemistry determination: A bacterial β-1,2-glucanase from P. gaetbulicola by monitoring the change in optical rotation <cite> | + | ;First stereochemistry determination: A bacterial β-1,2-glucanase from <i>P. gaetbulicola</i> by monitoring the change in optical rotation <cite>Nakajima2025</cite>. |
;First general base residue identification: not known. | ;First general base residue identification: not known. | ||
;First general acid residue identification: not known. | ;First general acid residue identification: not known. | ||
| − | ;First 3-D structure: A bacterial β-1,2-glucanase from <i>P. gaetbulicola</i> using the iodide single-wavelength anomalous diffraction phasing method. | + | ;First 3-D structure: A bacterial β-1,2-glucanase from <i>P. gaetbulicola</i> using the iodide single-wavelength anomalous diffraction phasing method <cite>Nakajima2025</cite>. |
== References == | == References == | ||
| Line 52: | Line 55: | ||
#Nakajima2025 pmid=40411428 | #Nakajima2025 pmid=40411428 | ||
#Tanaka2019 pmid=30926603 | #Tanaka2019 pmid=30926603 | ||
| − | #Abe2017 pmid= | + | #Tanaka2024 pmid=38300345 |
| + | #Abe2017 pmid=28270506 | ||
</biblio> | </biblio> | ||
Latest revision as of 22:48, 13 March 2026
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH194 | |
| Clan | GH-S |
| Mechanism | inverting |
| Active site residues | not known |
| CAZy DB link | |
| https://www.cazy.org/GH194.html | |
Substrate specificities
PgSGL3(H744_1c0222, KEGG) from Photobacterium gaetbulicola was characterized as reported in 2025 [1]. The enzyme specifically hydrolyzes β-1,2-glucan to produce β-1,2-glucooligosaccharides in an endolytic manner. PgSGL3 preferentially produces β-1,2-glucooctasaccharide at the initial stage of hydrolysis of β-1,2-glucan.
Kinetics and Mechanism
Hydrolysis of β-1,2-glucan by PgSGL3 suggests that the enzyme follows anomer-inverting mechanism [1]. Analysis of the change of the degree of optical rotation during hydrolysis of β-1,2-glucan and after addition of aqueous ammonia. Sharp decrease of the degree of optical rotation by aqueous ammonia is the same pattern as in the case of GH162 β-1,2-glucanase from Talaromyces funiculosus (TfSGL), an anomer-inverting enzyme [2].
Catalytic Residues
E214(PgSGL3) is the putative general acid as this residue is structurally well-superimposed with the general acid (E262) in GH162 TfSGL [2]. E214Q mutant shows drastic decrease in catalytic activity compared to the wild-type enzyme [1]. E214 is also conserved across other GH-S clan families including GH144, GH192, and GH193. In GH189, a family related to clan GH-S, this equivalent residue acts as a catalytic acid/base [3]
Similarly, D148 (PgSGL3) is a residue conserved spatially with several β-1,2-glucanases; GH144 (from Chitinophaga pinensis and Xanthomonas campestris pv. campestris), GH192 (from P. gaetbulicala), and GH193 (from Sanguibacter keddieii) [1, 4]. D148N mutant also shows drastically decreased activity against the wild-type enzyme. Mutational analysis alone is insufficient to definitively identify catalytic residues because a reaction mechanism of GH194 is atypical.
A plausible substrate binding mode of PgSGL3 can be obtained by superimposed with the complex structure of GH144 β-1,2-glucanase from X. campestris pv. campestris with β-1,2-glucoheptaose. However, no nucleophilic water is observed and no clear pathway for proton transfer from a nucleophilic water to a general base can be traced. It should be noted that the position of D148 (PgSGL3) does not correspond to that of the general base in GH162 TfSGL nor to the nucleophile in GH189 β-1,2-glucanotransferase [1, 2, 3], which suggests a difference in reaction mechanism between these families.
Three-dimensional structures
A ligand-free structure of PgSGL3 is available (PDB ID, 8XUK) [1]. PgSGL3 is composed of a single (α/α)6-barrel fold. The overall structure and the shape of catalytic pocket of PgSGL3 are similar to those of GH144 β-1,2-glucanases. The two candidate catalytic residues described above are well-superimposed with GH144 β-1,2-glucanases. Based on the similarity, GH194 is classified into clan GH-S, the same clan as GH144. Clan GH-S is composed of GH144, GH162, GH192, GH193, and GH194 which are distantly related families. Although GH189 is also a family distantly related to these families, GH189 is excluded from clan GH-S due to the difference in the reaction mechanism (GH189 enzymes follow anomer-retaining mechanism). These distantly related families including GH189 is called SGL clan [1]. The three residues labelled in red in the catalytic pocket (Fig. 1) are conserved within GH144, GH192 and GH193 families. The three conserved residues are considered as residues defining the SGL clan. Among the three residues, the glutamate residue is the candidate general acid described above.
Family Firsts
- First stereochemistry determination
- A bacterial β-1,2-glucanase from P. gaetbulicola by monitoring the change in optical rotation [1].
- First general base residue identification
- not known.
- First general acid residue identification
- not known.
- First 3-D structure
- A bacterial β-1,2-glucanase from P. gaetbulicola using the iodide single-wavelength anomalous diffraction phasing method [1].
References
- Nakajima M, Tanaka N, Motouchi S, Kobayashi K, Shimizu H, Abe K, Hosoyamada N, Abara N, Morimoto N, Hiramoto N, Nakata R, Takashima A, Hosoki M, Suzuki S, Shikano K, Fujimaru T, Imagawa S, Kawadai Y, Wang Z, Kitano Y, Nihira T, Nakai H, and Taguchi H. (2025). New glycoside hydrolase families of β-1,2-glucanases. Protein Sci. 2025;34(6):e70147. DOI:10.1002/pro.70147 |
- Tanaka N, Nakajima M, Narukawa-Nara M, Matsunaga H, Kamisuki S, Aramasa H, Takahashi Y, Sugimoto N, Abe K, Terada T, Miyanaga A, Yamashita T, Sugawara F, Kamakura T, Komba S, Nakai H, and Taguchi H. (2019). Identification, characterization, and structural analyses of a fungal endo-β-1,2-glucanase reveal a new glycoside hydrolase family. J Biol Chem. 2019;294(19):7942-7965. DOI:10.1074/jbc.RA118.007087 |
- Tanaka N, Saito R, Kobayashi K, Nakai H, Kamo S, Kuramochi K, Taguchi H, Nakajima M, and Masaike T. (2024). Functional and structural analysis of a cyclization domain in a cyclic β-1,2-glucan synthase. Appl Microbiol Biotechnol. 2024;108(1):187. DOI:10.1007/s00253-024-13013-9 |
- Abe K, Nakajima M, Yamashita T, Matsunaga H, Kamisuki S, Nihira T, Takahashi Y, Sugimoto N, Miyanaga A, Nakai H, Arakawa T, Fushinobu S, and Taguchi H. (2017). Biochemical and structural analyses of a bacterial endo-β-1,2-glucanase reveal a new glycoside hydrolase family. J Biol Chem. 2017;292(18):7487-7506. DOI:10.1074/jbc.M116.762724 |