CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Difference between revisions of "Carbohydrate Esterase Family 2"
Bobby Lamont (talk | contribs) |
Bobby Lamont (talk | contribs) |
||
| Line 12: | Line 12: | ||
|- | |- | ||
|'''Clan''' | |'''Clan''' | ||
| − | | | + | |α/β-hydrolase |
|- | |- | ||
|'''Mechanism''' | |'''Mechanism''' | ||
Revision as of 10:27, 16 November 2020
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
- Author: ^^^Bobby Lamont^^^
- Responsible Curators: ^^^Joel Weadge^^^ and ^^^Anthony Clarke^^^
| Carbohydrate Esterase Family CE2 | |
| Clan | α/β-hydrolase |
| Mechanism | Serine Hydrolase |
| Active site residues | Known |
| CAZy DB link | |
| https://www.cazy.org/CE2.html | |
Substrate specificities
All of the well characterized carbohydrate esterase family 2 enzymes have been shown to remove acetate groups from the synthetic molecule, 4-nitrophenyl acetate (4p-NP-Acc) [1, 2]. Contrary to typical acetyl xylan esterases, CE2 family members are shown to have a strong preference for the deacetylation of xylopyranosides at positions 3 and 4 instead of the typical deacetylation at position 2. CE2 family members were also shown to have significant preference for deacetylation of glucopyranosyl and mannopyranosyl residues at the 6-O position and significantly preferred deacetylation of glucopyranosyl and mannopyranosyl residues relative to the deacetylation of xylopyranosides. For these reasons CE2 family members are considered to be 6-de-O-acetylases [3].
Catalytic Residues
Most CE2 family members contain a catalytic dyad (Ser-His) as opposed to a catalytic triad that is typically found in esterases. CjCE2A is an exception and contains a Ser-His-Asp catalytic triad [1]. The structurally characterized, CtCE2, CjCE2B, and Est2A are examples of CE2 enzymes that contain catalytic dyads of conserved serine and histidine residues and lack the aspartate residue found in the triad. Without the support of an aspartate residue, the histidine residue of the catalytic dyads are supported by main-chain carbonyl groups provided by a backbone amino acid. The catalytic aspartate residue that would commonly complete the catalytic triad simply does not exist in many CE2 members. When CE2 enzymes contain a potential catalytic aspartate residue, often enough, there also exists a tryptophan residue that sits between the catalytic histidine and aspartate residues thereby prevent the aspartate residue from completing the triad. CE2 enzymes also possess an aromatic residue (either a tyrosine or a tryptophan residue) above their binding clefts that promote greater substrate specificity [1, 2]. Lastly, the oxyanion hole is comprised of the catalytic serine residue, a glycine, and an asparagine residue that appears to be invariant across the CE2 family and that of other related acetyl-esterases [1, 2].
Kinetics and Mechanism
The possession of an α/β hydrolase fold containing the catalytic serine nucleophile suggests that the reaction mechanism may proceed similar to other enzymes in the SGNH family. An example of a proposed reaction mechanism associated with the SGNH family of enzymes begins with the catalytic histidine residue acting as a base and increasing the nucleophilicity of the catalytic serine by extracting a proton from the hydroxyl group of the serine and leaving a negatively charged oxygen sidechain. The serine will attack the ester bond of the substrate, and stabilized by the residues of the oxyanion hole, a serine-substrate tetrahedral intermediate will form. The histidine acts as a general acid and donates a proton to the substrate causing its release and leaving an acetyl group attached to the serine. The histidine acts as a general base and influences a water molecule to attack the ester of the acetyl group attached to the catalytic serine generating a new tetrahedral intermediate state. The acetyl groups is released and the histidine will act as a general acid and donate a proton to back to the serine and restore the catalytic site to it’s original state [4].
The characterized enzymes were all tested using pNP-Ac, which showed kcat/KM of 2.01, 0.71, 0.38 and 3.13 s-1×µM-1 for Est2A, CtCE2, CjCE2A, and CjCE2B respectively. Est2A was also tested using p-nitrophenyl butyrate which gave a kcat/KM value of 2.33 x 10-3 s-1×µM-1 showing the significant decrease in catalytic efficiency as substrate size increased [2]. In order to test for positional specificity, the enzyme kinetics of CtCE2, CjCE2B, and CjCE2C were tested using 2-, 3-, and 4-O-acetyl p4-nitrophenyl β-D-xylopyranosides that showed increased kcat/KM values for the hydrolysis of the substrate at position 3 and 4 as opposed to position 2 [3]. Enzyme kinetic assays on birchwood xylan showed kcat/KM values of 7.33 x 10-5, 7.67 x 10-4, and 2.33 x 10-4 s-1xµM-1 for CtCE2, CjCE2A, and CjCE2B respectively. When the assay was performed on these same enzymes but with the use of glucomannan, the kcat/KM values were 9.67 x 10-4, 6.5 x 10-4, and 2.68 x 10-2 s-1xµM-1 for CtCE2, CjCE2A and CjCE2B respectively; thereby suggesting for these enzymes at least a clear substrate preference for glucomannan [1]
Three-dimensional structures
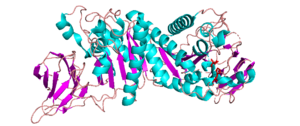
There are four reported structures for the CE2 family. These structures are all reported to be α/β-hydrolases and include Clostridium thermocellum’s CtCE2 (PDB ID 2WAO), Cellvibrio japonicus’ CjCE2A (PDB ID 2WAA) and CjCE2B (PDB ID 2W9X), and Butyrivibrio proteoclasticus’ Est2A (PDB ID 3U37). They contain an N-terminal β-sheet “jelly-roll” domain that acts as a carbohydrate binding domain (CBM) and is linked to a C-terminal domain that contains a α/β-hydrolase fold (SGNH-hydrolase motif) [1, 2]. This α/β-hydrolase fold consists of a three layered α/β stack composed of five beta strands arranged in parallel that form a central β-sheet, which is packed between α-helicies. In the case of CtCE2, CjCE2A, and CjCE2B, the sheet has 5 α-helices in total packed on each side [1], but Est2A, has 9 α-helices packing both sides of the β-sheet. The common structure of the N-terminal B-sheet “jelly-roll” domain across CE2 enzymes is comprised of two opposing β-sheets that have 4 and 5 β-strands, respectively [2].
The CE2 family members are typically monomeric, but there are some exceptions. Specifically, Est2A has been found to form tetramers that combine to make an overall octameric structure [1, 2]. The overall structure of CtCE2 also displayed that this domain is connected to the C-terminal end of a GH5 family cellulase protein, CtCel5C (PDB ID 4IM4), which make up a modular protein, called, CtCel5C-CE2. This protein is incorporated into cell-wall degrading cellulosome in C. thermocellum [1, 5].
Family Firsts
- First characterized
The first instance of CE2 family characterization was from the investigation of the BnaA, BnaB, and BnaC proteins that were discovered via cDNA library sequencing of Neocallimastix patriciarum [6]. BnaA and BnaC proteins exhibited acetyl xylan esterase ability. BnaB exhibited high sequence similarity to an uncharacterized C-terminal domain of the protein, CelE [7]. The uncharacterized domain of CelE which would later become known as, CtCE2 [1].
- First mechanistic insight
The catalytic dyad of Ser-His residues was confirmed by the arrangement of these residues in the crystal structures of CtCE2 (PDB ID 2WAO), CjCE2A (PDB ID 2WAA), and CjCE2B (PDB ID 2W9X) as well as sequence alignment analysis showing its invariance across the CE2 family [1].
- First 3-D structure
The first 3-D structures of CE2 family members, CtCE2, CjCE2A and CjCE2B were revealed in the same study that confirmed the catalytic mechanism of the family [1].
References
- Montanier C, Money VA, Pires VM, Flint JE, Pinheiro BA, Goyal A, Prates JA, Izumi A, Stålbrand H, Morland C, Cartmell A, Kolenova K, Topakas E, Dodson EJ, Bolam DN, Davies GJ, Fontes CM, and Gilbert HJ. (2009). The active site of a carbohydrate esterase displays divergent catalytic and noncatalytic binding functions. PLoS Biol. 2009;7(3):e71. DOI:10.1371/journal.pbio.1000071 |
- Till M, Goldstone DC, Attwood GT, Moon CD, Kelly WJ, and Arcus VL. (2013). Structure and function of an acetyl xylan esterase (Est2A) from the rumen bacterium Butyrivibrio proteoclasticus. Proteins. 2013;81(5):911-7. DOI:10.1002/prot.24254 |
- Topakas E, Kyriakopoulos S, Biely P, Hirsch J, Vafiadi C, and Christakopoulos P. (2010). Carbohydrate esterases of family 2 are 6-O-deacetylases. FEBS Lett. 2010;584(3):543-8. DOI:10.1016/j.febslet.2009.11.095 |
- Alalouf O, Balazs Y, Volkinshtein M, Grimpel Y, Shoham G, and Shoham Y. (2011). A new family of carbohydrate esterases is represented by a GDSL hydrolase/acetylxylan esterase from Geobacillus stearothermophilus. J Biol Chem. 2011;286(49):41993-42001. DOI:10.1074/jbc.M111.301051 |
- Bayer EA, Belaich JP, Shoham Y, and Lamed R. (2004). The cellulosomes: multienzyme machines for degradation of plant cell wall polysaccharides. Annu Rev Microbiol. 2004;58:521-54. DOI:10.1146/annurev.micro.57.030502.091022 |
- Dalrymple BP, Cybinski DH, Layton I, McSweeney CS, Xue GP, Swadling YJ, and Lowry JB. (1997). Three Neocallimastix patriciarum esterases associated with the degradation of complex polysaccharides are members of a new family of hydrolases. Microbiology (Reading). 1997;143 ( Pt 8):2605-2614. DOI:10.1099/00221287-143-8-2605 |
- Hall J, Hazlewood GP, Barker PJ, and Gilbert HJ. (1988). Conserved reiterated domains in Clostridium thermocellum endoglucanases are not essential for catalytic activity. Gene. 1988;69(1):29-38. DOI:10.1016/0378-1119(88)90375-7 |