CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Polysaccharide epimerases
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Margrethe Gaardlos^^^ and ^^^Anne Tondervik^^^
- Responsible Curator: ^^^Finn Aachmann^^^
Introduction
Classification
Mannuronan C5-epimerases
Substrate specificity
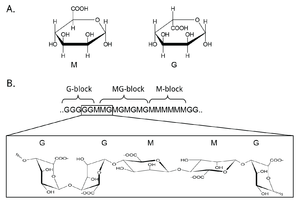
Mannuronan C5-epimerases are a group of enzymes that catalyze epimerization at the polymer-level of β-d-mannuronic acid residues (hereafter denoted M) into α-l-guluronic acid residues (hereafter denoted G) in alginate [1, 2, 3]. Alginate is an anionic polysaccharide made by brown seaweeds, some species of red algae, and the gram-negative bacterial genera Pseudomonas and Azotobacter [4, 5, 6, 7, 8]. The function of alginate in the different organisms are various, and related to structure, protection and surface adhesion [9, 10, 11, 12]. Alginate is a copolymer of the two 1-4 linked epimers [13, 14, 15], and by changing the composition of the two monomers the epimerases fine-tune the properties of the polymer [16].
At first, alginate is made as a homopolymer of M in the cell. Epimerases then convert some of the M residues in the polymer into G-residues [3, 17, 18]. This epimerization is not random and creates block structures of M, G or alternating MG [19, 20]. Alginate residues that are oxidized or acetylated are not substrates for the epimerases, and acetylation of alginate could be a way to control epimerization in nature [21, 22].
Mannuronan C5-epimerases exist both in algae and in bacteria [1, 23]. Gene analyses propose as many as 31 different genes encoding putative mannuronan C-5 epimerases in the brown algae Ectocarpus [24]. However, the algal epimerases are difficult to express and it is the bacterial enzymes that have been studied most extensively [24, 25]. Two categories of bacterial mannuronan C-5-epimerases have been described: the periplasmic AlgG and the extracellular and calcium dependent AlgE. AlgG creates single G residues in stretches of mannuronan, while the AlgE enzymes are processive and create MG-blocks and G-blocks. Pseudomonas is only known to produce AlgG [18, 26, 27], while A. vinelandii contains seven active AlgE enzymes in addition to AlgG [28, 29, 30, 31]. A mutant strain of P. fluorescens without the algG gene creates pure mannuronan \autocite{Gimmestad2003}. This strain can be used to produce unepimerized substrate, which is useful for the study of the epimerization reaction. Methods for studying this are discussed in Section \ref{sec:catmec}.
Product profiles
Main section 2
Whatevs...
References
- Haug A and Larsen B. (1969). Biosynthesis of alginate. Epimerisation of D-mannuronic to L-guluronic acid residues in the polymer chain. Biochim Biophys Acta. 1969;192(3):557-9. DOI:10.1016/0304-4165(69)90414-0 |
- Larsen B and Haug A. (1971). Biosynthesis of alginate. 1. Composition and structure of alginate produced by Azotobacter vinelandii (Lipman). Carbohydr Res. 1971;17(2):287-96. DOI:10.1016/s0008-6215(00)82536-7 |
- Haug A and Larsen B. (1971). Biosynthesis of alginate. II. Polymannuronic acid C-5-epimerase from Azotobacter vinelandii (Lipman). Carbohydr Res. 1971;17(2):297-308. DOI:10.1016/s0008-6215(00)82537-9 |
-
Stanford, Edw C C. (1883) On algin: a new substance obtained from some of the commoner species of marine algae. R. Anderson. NLM ID: 101217546
-
Gorin, P. A. J. and Spencer, J. F. T. (1966) Exocellular alginic acid from Azotobacter vinelandii. Canadian Journal of Chemistry vol. 44, no. 9., pp. 993-998. [1]
- Linker A and Jones RS. (1966). A new polysaccharide resembling alginic acid isolated from pseudomonads. J Biol Chem. 1966;241(16):3845-51. | Google Books | Open Library
- Govan JR, Fyfe JA, and Jarman TR. (1981). Isolation of alginate-producing mutants of Pseudomonas fluorescens, Pseudomonas putida and Pseudomonas mendocina. J Gen Microbiol. 1981;125(1):217-20. DOI:10.1099/00221287-125-1-217 |
-
Okazaki, M., K. and Furuya, K. Tsukayam and K. Nisizawa. (1982) Isolation and Identification of Alginic Acid from a Calcareous Red Alga Serraticardia maxima. Botanica Marina, vol. 25, no. 3., pp. 123-131. [1]
-
Painter, Terence J. (1983) Chapter 4 - Algal Polysaccharides. Edited by Gerald O. Aspinall. The Polysaccharides. New York: Academic Press. [1]
- Campos M, Martínez-Salazar JM, Lloret L, Moreno S, Núñez C, Espín G, and Soberón-Chávez G. (1996). Characterization of the gene coding for GDP-mannose dehydrogenase (algD) from Azotobacter vinelandii. J Bacteriol. 1996;178(7):1793-9. DOI:10.1128/jb.178.7.1793-1799.1996 |
- Pier GB, Coleman F, Grout M, Franklin M, and Ohman DE. (2001). Role of alginate O acetylation in resistance of mucoid Pseudomonas aeruginosa to opsonic phagocytosis. Infect Immun. 2001;69(3):1895-901. DOI:10.1128/IAI.69.3.1895-1901.2001 |
- Harmsen M, Yang L, Pamp SJ, and Tolker-Nielsen T. (2010). An update on Pseudomonas aeruginosa biofilm formation, tolerance, and dispersal. FEMS Immunol Med Microbiol. 2010;59(3):253-68. DOI:10.1111/j.1574-695X.2010.00690.x |
-
Hirst, E. L. and Jones, J. K. N and Jones, Winifred Osman. (1939) 389. The structure of alginic acid. Part I [in en]. Journal of the Chemical Society, The Royal Society of Chemistry. Vol. 0, pp. 1880–1885. [1]
- FISCHER FG and DORFEL H. (1955). [Polyuronic acids in brown algae]. Hoppe Seylers Z Physiol Chem. 1955;302(4-6):186-203. | Google Books | Open Library
-
Drummond, D W and Hirst, E L and Percival, Elizabeth. (1962) 232. The constitution of alginic acid. Journal of the Chemical Society, The Royal Society of Chemistry. Vol. 0, pp. 1208–1216. [1]
- Ertesvåg H, Høidal HK, Schjerven H, Svanem BI, and Valla S. (1999). Mannuronan C-5-epimerases and their application for in vitro and in vivo design of new alginates useful in biotechnology. Metab Eng. 1999;1(3):262-9. DOI:10.1006/mben.1999.0130 |
- Lin TY and Hassid WZ. (1966). Pathway of algnic acid synthesis in the marine brown alga, Fucus gardneri Silva. J Biol Chem. 1966;241(22):5284-97. | Google Books | Open Library
- Franklin MJ, Chitnis CE, Gacesa P, Sonesson A, White DC, and Ohman DE. (1994). Pseudomonas aeruginosa AlgG is a polymer level alginate C5-mannuronan epimerase. J Bacteriol. 1994;176(7):1821-30. DOI:10.1128/jb.176.7.1821-1830.1994 |
-
Haug, Arne and Larsen, Bjørn and Smidsrød, Olav. (1966) A study on the constitution of alginic acidby partial acid hydrolysis. Acta Chemica Scandinavica, vol. 5 (July), pp. 271–277. [1]
-
Haug, Arne and Larsen, Bjørn and Smidsrød, Olav. (1967) Studies on the Sequence of Uronic Acid Residues in Alginic Acid. Acta Chemica Scandinavica, vol. 21, pp. 691–794. [1]
-
Skjåk-Bræk, Gudmund and Larsen, Bjørn and Grasdalen, Hans. (1985) The role of O-acetyl groupsin the biosynthesis of alginate by Azotobacter vinelandii. Carbohydrate Research, vol. 145, no. 1, pp. 169–174. [1]
-
Kristiansen, Kåre A and Schirmer, Bjørn C and Aachmann, Finn L. and Skjåk-Bræk, Gudmund and Draget, Kurt I. and Christensen, Bjørn E. (2009) Novel alginates prepared by independent control of chain stiff-ness and distribution of G-residues: Structure and gelling properties. Carbohydrate Polymers, vol. 77, no.4, pp. 725–735. [1]
- Madgwick J, Haug A, and Larsen B. (1973). Polymannuronic acid 5-epimerase from the marine alga Pelvetia canaliculata (L.) Dcne. et Thur. Acta Chem Scand. 1973;27(9):3592-4. DOI:10.3891/acta.chem.scand.27-3592 |
- Fischl R, Bertelsen K, Gaillard F, Coelho S, Michel G, Klinger M, Boyen C, Czjzek M, and Hervé C. (2016). The cell-wall active mannuronan C5-epimerases in the model brown alga Ectocarpus: From gene context to recombinant protein. Glycobiology. 2016;26(9):973-983. DOI:10.1093/glycob/cww040 |
- Nyvall P, Corre E, Boisset C, Barbeyron T, Rousvoal S, Scornet D, Kloareg B, and Boyen C. (2003). Characterization of mannuronan C-5-epimerase genes from the brown alga Laminaria digitata. Plant Physiol. 2003;133(2):726-35. DOI:10.1104/pp.103.025981 |
- Chitnis CE and Ohman DE. (1990). Cloning of Pseudomonas aeruginosa algG, which controls alginate structure. J Bacteriol. 1990;172(6):2894-900. DOI:10.1128/jb.172.6.2894-2900.1990 |
- Morea A, Mathee K, Franklin MJ, Giacomini A, O'Regan M, and Ohman DE. (2001). Characterization of algG encoding C5-epimerase in the alginate biosynthetic gene cluster of Pseudomonas fluorescens. Gene. 2001;278(1-2):107-14. DOI:10.1016/s0378-1119(01)00685-0 |
- Ertesvåg H, Doseth B, Larsen B, Skjåk-Braek G, and Valla S. (1994). Cloning and expression of an Azotobacter vinelandii mannuronan C-5-epimerase gene. J Bacteriol. 1994;176(10):2846-53. DOI:10.1128/jb.176.10.2846-2853.1994 |
- Ertesvåg H, Høidal HK, Hals IK, Rian A, Doseth B, and Valla S. (1995). A family of modular type mannuronan C-5-epimerase genes controls alginate structure in Azotobacter vinelandii. Mol Microbiol. 1995;16(4):719-31. DOI:10.1111/j.1365-2958.1995.tb02433.x |
- Rehm BH, Ertesvåg H, and Valla S. (1996). A new Azotobacter vinelandii mannuronan C-5-epimerase gene (algG) is part of an alg gene cluster physically organized in a manner similar to that in Pseudomonas aeruginosa. J Bacteriol. 1996;178(20):5884-9. DOI:10.1128/jb.178.20.5884-5889.1996 |
- Svanem BI, Skjåk-Braek G, Ertesvåg H, and Valla S. (1999). Cloning and expression of three new Aazotobacter vinelandii genes closely related to a previously described gene family encoding mannuronan C-5-epimerases. J Bacteriol. 1999;181(1):68-77. DOI:10.1128/JB.181.1.68-77.1999 |
- Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, and Henrissat B. (2009). The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res. 2009;37(Database issue):D233-8. DOI:10.1093/nar/gkn663 |
-
Davies, G.J. and Sinnott, M.L. (2008) Sorting the diverse: the sequence-based classifications of carbohydrate-active enzymes. The Biochemist, vol. 30, no. 4., pp. 26-32. Download PDF version.