CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Carbohydrate Esterase Family 7
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Carbohydrate Esterase Family CE7 | |
| Fold | (α/β/α)-Sandwich |
| Mechanism | Serine Hydrolase |
| Active site residues | Known, Catalytic Triad |
| CAZy DB link | |
| https://www.cazy.org/CE7.html | |
Substrate specificities
Carbohydrate Esterase Family 7 currently contains enzymes classified as acetyl xylan esterases (AXEs) and cephalosporin-C deacetylases [1]. One exception is the AXE from Thermotoga martitima that lacks detectable activity towards acetylated xylan, but rather, shorter acetylated xylo-oligomers [2]. A common feature of the oligomeric enzymes belonging to this family is that they usually serve as multifunctional deacetylases and are active on a broad range of small substrates with relatively low acetyl-positional specificity [2, 3, 4, 5]. Some examples include xylose tetraacetate, glucose pentaacetate, α-napthyl acetate, 4-methylumbelliferyl acetate and p-nitrophenyl acetate (pNP-acetate) [6]. Substrate binding is proposed to be largely mediated through non-specific hydrophobic interactions with surface amino acid residues [5].
Catalytic Residues
Members of this family use the catalytic triad of serine, histidine and aspartate [7]. The aspartate promotes the amphoteric nature of the histidine residue so that it can abstract a proton from the serine residue to render the serine nucleophilic [2]. The Gly-X-Ser-X-Gly motif, containing the nucleophilic serine, is common amongst the broad α/β-hydrolase family to which the CE7 enzymes belong [8]. The glycine residues near the catalytic serine are important to prevent any steric hindrance that may occur from amino acids with larger side chains and form the basis for the nucleophilic “elbow” turn motif that is present in these enzymes [2].
Kinetics and Mechanism
The CE7 family of esterases catalyze the removal of O-acetyl groups from cephalosporin-C, xylan and xylo-oligosaccharides [5, 7]. Members of this family present their active site residues in typical positions observed for α/β-hydrolases [9]. The catalytic triad residues responsible for de-O-acetylation are found on loops near the C-terminus of the β-strands 5, 7 and 8 for the catalytic serine, aspartate and histidine, respectively [10]. Similar to other α/β-hydrolases, the CE7 enzymes follow a mechanism where the serine is deprotonated by the histidine imidazole side chain; thereby allowing for nucleophilic attack of the serine on the incoming carbonyl of the substrate’s acetyl group [9]. The result is a transient tetrahedral oxyanion intermediate, which is stabilized by the backbone amide group of a highly conserved glutamate residue (found in a signature RGQ motif adjacent to the nucleophilic serine at the end of β-strand 4) and a less conserved tyrosine that together form the oxyanion hole of the enzyme [2, 5]. The tetrahedral intermediate then collapses resulting in an acetyl-enzyme intermediate and release of the de-O-acetylated product. A water molecule entering the active site is deprotonated by histidine and then nucleophilic attack by water on the acetyl-enzyme takes place to liberate acetate and the free enzyme [9]. A recent study on the CE7 AXE (TM0077) from T. martitima provided further support into the catalytic mechanism [2]. TM0077 was co-crystallized with PMSF and paraoxan (PDB ID 3M82 and PDB ID 3M83, respectively), two known serine protease inhibitors, that led to a 110° rotation of the catalytic serine and the formation of a covalent tetrahedral intermediate that was unable to collapse back to the starting materials (TM0077 and PMSF/paraoxan) [2]. Trapping the tetrahedral intermediate is strong evidence that CE7 enzymes proceed by the proposed mechanism.
Kinetic parameters and specific activities have been determined for some members of CE7. CAH from Bacillus subtilis, AXE from Paenibacillus sp. R4 and AXE (TM0077) from T. maritima were assayed for their activity against pNP-acetate and were found to have KM values of 0.29 +/- 0.01, 0.15 +/- 0.01 and 0.19 +/- 0.03 mM, respectively, and kcat/KM values of 260, 360 and 310 s-1mM-1, respectively [2, 5, 11, 12]. There has been discrepancy regarding the ability of these CE7 enzymes to utilize acetylated xylan as a substrate. While a Bacillus pumilus AXE has demonstrated a specific activity against acetylated xylan of 41 +/- 8 U/mg, T. maritima's AXE had no demonstrated activity and the acetyl xylan esterase, Axe1, from Thermoanerobacterium saccharolyticum strain JW/SL-YS485 had no activity in one study and 5.2 U/mg (8-fold lower than the B. pumilus enzyme) in another study [2, 4, 13, 14]. In light of the apparent low/no activity against acetylated xylan, the enzymes from T. maritima and T. saccharolyticum were assayed against smaller substrates, where the T. maritima AXE (TM0077) had high activity against glucose pentaacetate with a kcat of 2900 s-1 and Axe1 from T. saccharolyticum had high activity against xylose tetraacetate with separately reported specific activities of 210 and 740 U/mg [2, 4, 14]. Lastly, CAH from B. subtilis has been kinetically characterized with regards to activity towards cephalosporin-C with a KM of 24 mM and a kcat/KM of 7.0 s-1mM-1, which has supported its naming as a cephalosporin-C deacetylase [11].
Three-dimensional structures
Several structures have been reported for the CE7 family. These include the AXE from B. pumilus (PDB ID 2XLB), cephalosporin-C deacetylase (CAH) from B. subtilis strain 168 (PDB ID 1ODS), the AXE from Paenibacillus sp. R4 (PDB ID 6AGQ), acetyl xylan esterase 1 (Axe1) from T. saccharolyticum strain JW/SL-YS485 (PDB ID 3FCY), the AXE (TM0077) from T. maritima (PDB ID 1VLQ) and, lastly, an unclassified protein named Axe1-NaM1 (PDB ID 6FKX).
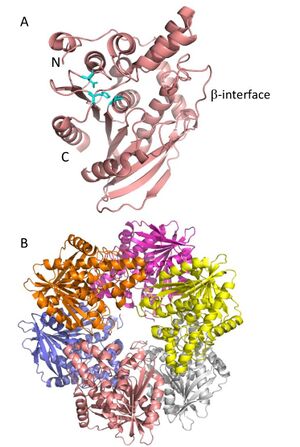
Enzymes in the CE7 family contain high levels of multimerization, with the quaternary structure adopting a dimer of trimer hexamer structure[5]. For each of the six resolved structures from the CE7 family, individual subunits adopt a classic α/β-hydrolase fold that consists of a central eight-stranded β-sheet surrounded by α-helices on both sides (See Fig. 1A) [2]. Together, the multimerized subunits adopt a donut-like shape that results in the formation of a tunnel leading to the centre of the protein (See Fig. 1B). Each of the active sites from the individual multimers are arranged so that they face into the centre of the tunnel [5].
Family Firsts
- First Characterized
- The first cephalosporin-C deacetylase (CAH) of this family to be characterized was from B. subtilis strain 168 in 1975 [15]. CAH is able to deacetylate cephalosporin-C, as well as cephalosporin-C derivatives (eg., cephalosporanic acid and cephalothin) at the C-3 position [15]. The first acetyl xylan esterase to be characterized was Axe1, from the anaerobic thermophilic bacterium T. saccharolyticum strain JW/SL-YS485 in 1995 [14]. When in the presence of acetylated xylan, Axe1 showed esterase activity that liberated acetate, leading to its name [14]. However, Axe1 has also demonstrated activity against other acetylated sugars [14].
- First Mechanistic Insight
- The first mechanistic insight into the CE7 family came from the CAH enzyme from B. subtilis in 1994 when Takimoto and coworkers tested the effects of various chemicals (eg., EDTA, PMSF, diisopropyl flurophosphate) on CAH activity and concluded that serine is likely important for catalysis [11]. Almost 10 years later in 2003, Vincent and colleagues resolved the crystal structure of CAH, confirming that serine was present in the active site and that an aspartate and histidine completed the catalytic triad [5]. More recently, co-crystallization of the AXE (TM0077) from T. martitima with PMSF and paraoxan (PDB ID 3M82 and PDB ID 3M83, respectively), two known serine protease inhibitors, led to trapping of the tetrahedral intermediate; thereby providing further support for the serine-histidine-aspartate mediated mechanism described for this family.
- First 3-D Structure
- The first resolved structure in this family was the cephalosporin-C deacetylase, CAH (PDB ID 1ODS) from B. subtilis strain 168 in 2003 [5]. The quaternary structure consisted of a trimer of dimers to make up a donut-shaped homohexamer with 32kDa subunits that each adopt the α/β hydrolase fold (See Fig. 1) [5]. The hydrophobic core containing the six active site centers is thought to help exclude large substrates[5]. In order for the hexamer to form, there are α-helix interactions as well as a β-strand-like interface associating between monomers (See Fig. 1A) [5, 9].
References
-
Nakamura, AM, Nascimento, AS and Polikarpov, I. (2017) Structural diversity of carbohydrate esterases. Biotechnol. Res. Innov., vol. 1, pp. 35-51. [1].
- Levisson M, Han GW, Deller MC, Xu Q, Biely P, Hendriks S, Ten Eyck LF, Flensburg C, Roversi P, Miller MD, McMullan D, von Delft F, Kreusch A, Deacon AM, van der Oost J, Lesley SA, Elsliger MA, Kengen SW, and Wilson IA. (2012). Functional and structural characterization of a thermostable acetyl esterase from Thermotoga maritima. Proteins. 2012;80(6):1545-59. DOI:10.1002/prot.24041 |
- Krastanova I, Guarnaccia C, Zahariev S, Degrassi G, and Lamba D. (2005). Heterologous expression, purification, crystallization, X-ray analysis and phasing of the acetyl xylan esterase from Bacillus pumilus. Biochim Biophys Acta. 2005;1748(2):222-30. DOI:10.1016/j.bbapap.2005.01.003 |
- Lorenz WW and Wiegel J. (1997). Isolation, analysis, and expression of two genes from Thermoanaerobacterium sp. strain JW/SL YS485: a beta-xylosidase and a novel acetyl xylan esterase with cephalosporin C deacetylase activity. J Bacteriol. 1997;179(17):5436-41. DOI:10.1128/jb.179.17.5436-5441.1997 |
- Vincent F, Charnock SJ, Verschueren KH, Turkenburg JP, Scott DJ, Offen WA, Roberts S, Pell G, Gilbert HJ, Davies GJ, and Brannigan JA. (2003). Multifunctional xylooligosaccharide/cephalosporin C deacetylase revealed by the hexameric structure of the Bacillus subtilis enzyme at 1.9A resolution. J Mol Biol. 2003;330(3):593-606. DOI:10.1016/s0022-2836(03)00632-6 |
- Degrassi G, Kojic M, Ljubijankic G, and Venturi V. (2000). The acetyl xylan esterase of Bacillus pumilus belongs to a family of esterases with broad substrate specificity. Microbiology (Reading). 2000;146 ( Pt 7):1585-1591. DOI:10.1099/00221287-146-7-1585 |
- Biely P (2012). Microbial carbohydrate esterases deacetylating plant polysaccharides. Biotechnol Adv. 2012;30(6):1575-88. DOI:10.1016/j.biotechadv.2012.04.010 |
- Mitsushima K, Takimoto A, Sonoyama T, and Yagi S. (1995). Gene cloning, nucleotide sequence, and expression of a cephalosporin-C deacetylase from Bacillus subtilis. Appl Environ Microbiol. 1995;61(6):2224-9. DOI:10.1128/aem.61.6.2224-2229.1995 |
- Sista Kameshwar AK and Qin W. (2018). Understanding the structural and functional properties of carbohydrate esterases with a special focus on hemicellulose deacetylating acetyl xylan esterases. Mycology. 2018;9(4):273-295. DOI:10.1080/21501203.2018.1492979 |
- Ollis DL, Cheah E, Cygler M, Dijkstra B, Frolow F, Franken SM, Harel M, Remington SJ, Silman I, and Schrag J. (1992). The alpha/beta hydrolase fold. Protein Eng. 1992;5(3):197-211. DOI:10.1093/protein/5.3.197 |
-
Takimoto, A, Mitsushima, K, Yagi, S. and Sonoyama, T. (1994) Purification, Characterization and Partial Amino Acid Sequences of a Novel Cephalosporin-C Deacetylase from Bacillus subtilis. J. Fermentation Bioeng., vol. 77, no. 1, pp. 17-22.
- Park SH, Yoo W, Lee CW, Jeong CS, Shin SC, Kim HW, Park H, Kim KK, Kim TD, and Lee JH. (2018). Crystal structure and functional characterization of a cold-active acetyl xylan esterase (PbAcE) from psychrophilic soil microbe Paenibacillus sp. PLoS One. 2018;13(10):e0206260. DOI:10.1371/journal.pone.0206260 |
- Degrassi G, Okeke BC, Bruschi CV, and Venturi V. (1998). Purification and characterization of an acetyl xylan esterase from Bacillus pumilus. Appl Environ Microbiol. 1998;64(2):789-92. DOI:10.1128/AEM.64.2.789-792.1998 |
- Shao W and Wiegel J. (1995). Purification and characterization of two thermostable acetyl xylan esterases from Thermoanaerobacterium sp. strain JW/SL-YS485. Appl Environ Microbiol. 1995;61(2):729-33. DOI:10.1128/aem.61.2.729-733.1995 |
- Abbott BJ and Fukuda D. (1975). Physical properties and kinetic behavior of a cephalosporin acetylesterase produced by Bacillus subtilis. Appl Microbiol. 1975;30(3):413-9. DOI:10.1128/am.30.3.413-419.1975 |