CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 102
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH102 | |
| Clan | none |
| Mechanism | retaining |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/GH102.html | |
Substrate specificities
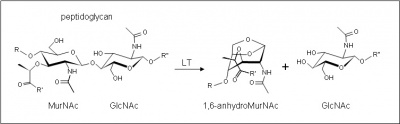
The glycoside hydrolases of this family are lytic transglycosylases (also referred to as peptidoglycan lyases) of bacterial origin and they constitute family 2 of the classification scheme of Blackburn and Clarke [1]. The prototype for this family is membrane-bound lytic transglycosylase A (MltA) from Escherichia coli. These enzymes cleave the β-1,4-linkage between N-acetylmuramoyl and N-acetylglucosaminyl residues in peptidoglycan (Figure 1), but unlike the lytic transglycosylases of other families, they are active on peptidoglycan fragments lacking their stem peptides [2]. No other activities have been observed.
Kinetics and Mechanism
The lytic transglycosidases, strictly speaking, are retaining enzymes. They are not hydrolases but rather catalyse an intramolecular glycosyl transferase reaction onto the C-6 hydroxyl group of the muramoyl residue leading to the generation of a terminal 1,6-anhydromuramoyl product (Figure 1) thus lacking a reducing end [3]. No detailed analyses involving both steady state and pre-steady state kinetic studies have been reported, but the Michaelis Menten (KM and Vmax) parameters have been estimated for E. coli MltA acting on insoluble peptidoglycan sacculi [2].
Catalytic Residues
As with other lytic transglycosylases (families GH23, GH103, and GH104), the GH102 enzymes are thought to possess a single catalytic general acid/base residue. This residue in E. coli MltA has been identified as Asp308 and, indeed, its replacement with Ala abolishes catalytic activity [4].
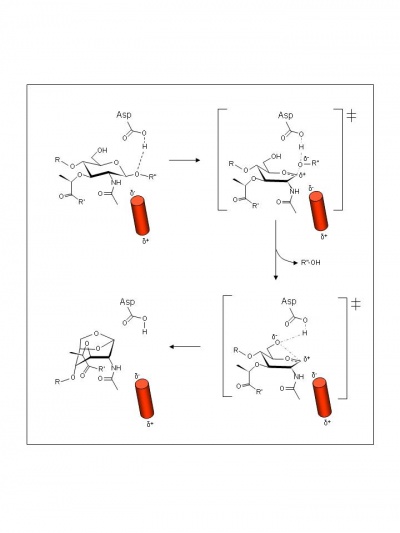
The mechanism of action of the family GH102 enzymes has yet to be proven experimentally, but examination of crystal structures of E. coli MltA complexed with chitohexoase has led to the proposal of a novel mechanism of action involving the stabilization of the putative intermediate oxocarbenium ion by an alpha-helix dipole [5]. Thus, the catalytic Asp308 is proposed to serve initially as an acid catalyst to donate a proton to the glycosidic oxygen of the linkage to be cleaved leading to the formation of an intermediate with oxocarbenium ion character (Figure 2). In the absence of an anion/nucleophile in close proximity, this oxocarbenium intermediate is proposed to be stabilized by the negatively-charged side of the dipole that would exist at the C-terminal end of an alpha helix positioned just below the -1 binding subsite. This would be followed by abstraction of the C-6 hydroxyl proton of the intermediate involving Asp308 which now serves as the base catalyst leading to nucleophilic attack and the formation of 1,6-anhydromuramic acid product.
Three-dimensional structures
Three-dimensional structures are available for several Family GH103 enzymes, the first solved being that of E. coli MltA [4]. Unlike the other lytic transglycosylases of families GH23, GH103, and GH104 which possesses the well characterized α+β “lysozyme fold,” these enzymes have a unique structure consisting of two domains. One has a double-ψ β-barrel fold similar to the catalytic domain of the family GH45 endoglucanase V from Humicola insolens [6]. The second and smaller domain has a β-barrel fold topology. The large groove between the two domains serves as the active site cleft.
Family Firsts
- First identification of lytic transglycosylase
- MltA from E. coli [2].
- First general acid/base residue identification
- Inferred by X-ray crystallography of E. coli MltA [4].
- First 3-D structure
- E. coli MltA [4].
- First identification as a lipoprotein
- E. coli MltA [7].
- First identification of localization to outer membrane
- E. coli MltA [7].
- Frist demonstration of molecular interactions between GH102 enzymes and penicillin-binding proteins
- E. coli MltA [8].
References
- Blackburn NT and Clarke AJ. (2001). Identification of four families of peptidoglycan lytic transglycosylases. J Mol Evol. 2001;52(1):78-84. DOI:10.1007/s002390010136 |
- Ursinus A and Höltje JV. (1994). Purification and properties of a membrane-bound lytic transglycosylase from Escherichia coli. J Bacteriol. 1994;176(2):338-43. DOI:10.1128/jb.176.2.338-343.1994 |
- Höltje JV, Mirelman D, Sharon N, and Schwarz U. (1975). Novel type of murein transglycosylase in Escherichia coli. J Bacteriol. 1975;124(3):1067-76. DOI:10.1128/jb.124.3.1067-1076.1975 |
- van Straaten KE, Dijkstra BW, Vollmer W, and Thunnissen AM. (2005). Crystal structure of MltA from Escherichia coli reveals a unique lytic transglycosylase fold. J Mol Biol. 2005;352(5):1068-80. DOI:10.1016/j.jmb.2005.07.067 |
- van Straaten KE, Barends TR, Dijkstra BW, and Thunnissen AM. (2007). Structure of Escherichia coli Lytic transglycosylase MltA with bound chitohexaose: implications for peptidoglycan binding and cleavage. J Biol Chem. 2007;282(29):21197-205. DOI:10.1074/jbc.M701818200 |
- Davies GJ, Dodson GG, Hubbard RE, Tolley SP, Dauter Z, Wilson KS, Hjort C, Mikkelsen JM, Rasmussen G, and Schülein M. (1993). Structure and function of endoglucanase V. Nature. 1993;365(6444):362-4. DOI:10.1038/365362a0 |
- Lommatzsch J, Templin MF, Kraft AR, Vollmer W, and Höltje JV. (1997). Outer membrane localization of murein hydrolases: MltA, a third lipoprotein lytic transglycosylase in Escherichia coli. J Bacteriol. 1997;179(17):5465-70. DOI:10.1128/jb.179.17.5465-5470.1997 |
- Vollmer W, von Rechenberg M, and Höltje JV. (1999). Demonstration of molecular interactions between the murein polymerase PBP1B, the lytic transglycosylase MltA, and the scaffolding protein MipA of Escherichia coli. J Biol Chem. 1999;274(10):6726-34. DOI:10.1074/jbc.274.10.6726 |