CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 186
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH186 | |
| Clan | None |
| Mechanism | Inverting |
| Active site residues | Asp |
| CAZy DB link | |
| https://www.cazy.org/GH186.html | |
Substrate specificities
The defining member of GH186, a β-1,2-glucanase from Escherichia coli (EcOpgD) was identified, characterized and structurally analyzed as reported in 2023 [1]. Subsequently, GH186 homolog from Xanthomonas campestris pv. campestris (XccOpgD) was found to be an anomer-inverting transglycosylase which specifically generated α-1,6-glucosidic bonds from β-1,2-glucan donors [2]. EcOpgD and XccOpgD are specific toward β-1,2-glucan and the amino acid residues for recognizing β-1,2-glucan at common subsites between EcOpgD and XccOpgD (subsite –7 to +6) are highly conserved in GH186 [1, 2]. However, the reaction types of EcOpgD and XccOpgD are different from each other [1, 2]. EcOpgD is a β-1,2-glucanase that preferentially generates β-1,2-glucooligosaccharides (Sopns, where n indicates a degree of polymerization, DP) with DPs of 6 and 7 from linear β-1,2-glucan [1]. Final products produced by EcOpgD are Sop6–10, indicating that EcOgpD hydrolyzes Sopns with DPs of 11 and higher [1]. XccOpgD generates only α-1,6-cyclized β-1,2-glucohexadecaose from linear β-1,2-glucan [2]. Almost all family members are found in Pseudomonadota, particularly gamma proteobacteria.
Kinetics and Mechanism
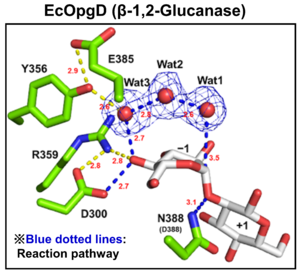
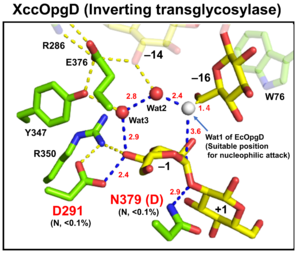
Optical rotation and NMR analyses indicated that EcOpgD and XccOpgD use anomer-inverting mechanisms [1, 2]. X-ray structural analysis (see below) and mutational analysis suggest that D388 in EcOpgD and the equivalent residue in XccOpgD, D379, directly protonate the scissile glycosidic bond as general acids (Figure 1, 2) [1, 2]. These analyses also suggested that D300 in EcOpgD and the equivalent residue in XccOpgD, D291, act as as general bases to activate the nucleophile via a chain comprising the 4-hydroxy group of the Glc moiety at subsite –1 and two water molecules [1, 2]. In EcOpgD (glucanase), the nucleophile is a water molecule and in XccOpgD (transglycosylase) the nucleophile is the 6-hydroxy group of the terminal Glc moiety at position –16 in the chain [1, 2]. It should be noted that transglycosylation is more typical of anomer-retaining GHs, which use a covalent glycosyl-enzyme intermediate, further highlighting the uniqueness of these GH186 members.
The difference in nucleophiles among GH186 family is probably due to how the nucleophile and the Grotthuss proton relay are stabilized. While W441, which is important for stabilizing nucleophilic water in EcOpgD, is not conserved in GH186, W76, which is important for stacking with the acceptor Glc moiety in XccOpgD is broadly conserved (but not in the clade of EcOpgD) (Figure 2) [1, 2]. Therefore, GH186 seems to be fundamentally an anomer-inverting transglycosylase family. In addition, the Grotthuss proton relay is sequestered (stabilized) not by amino acid sequence but by positioning of the acceptor substrate in XccOpgD, with the result that a water molecule is not suitable as a nucleophile for efficient Grotthuss proton relay. This explains why XccOpgD is a specific transglycosylase [2].
Catalytic Residues
As described above, the identities of the catalytic residues were indicated by X-ray crystallography and supported by site-directed mutagenesis. The general acid and general base of EcOpgD are D388 and D300, respectively, and the equivalent catalytic residues of XccOpgD are D379 and D291, respectively) [1, 2].
Three-dimensional structures
The ligand-free structure of OpgG from E. coli (EcOpgG) was determined at 2.4 Å resolution (PDB 1TXK) [3]. The ligand-free structure of EcOpgD was determined at 2.95 Å resolution (PDB 8IOX) [1]. Michaelis complexes of EcOpgD (D388N, co-crystal), EcOpgG (D361N, soaking) and XccOpgD (D379N, soaking) with β-1,2-glucan were determined at 2.06, 1.81, and 2.25 Å resolutions, respectively (PDB 8IP1, PDB 8IP2, PDB 8X18) [1, 2]. Notably, GH186 is strucuturally unique, i.e. lacks 3-D structural homologs, among all of the GHs families known at the time [1].
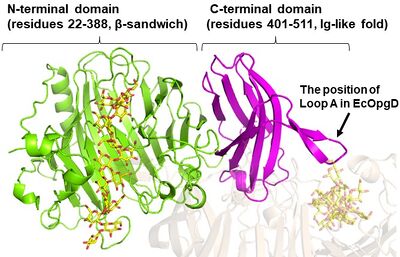
EcOpgG consists of an N-terminal domain (residues 22–388, β-sandwich) and a C-terminal domain (residues 401–511, Ig-like fold) (Figure 3). The two domains are connected with one turn of a 310 helix [1, 3]. The loop region (residues 409-425, Loop A below) in the C-terminal domain of the ligand-free structure organizes into β-strands in the Michaelis complex structure. In the Michaelis complex, the β-strands extend toward the catalytic center of another chain in the dimer to cover the proton transfer pathway from a nucleophile to the general base catalyst [1]. However, the sequence of Loop A is diversified in GH186 family. Indeed, Loop A in EcOpgD sequesters the proton transfer pathway from the solvent, while that of EcOpgG does not completely, which is consistent with the drastically reduced hydrolytic activity of EcOpgG compared with EcOpgD [1]. In addition, the corresponding Loop A of XccOpgD is too short to reach the catalytic center, which makes space to accommodate a β-1,2-glucooligosaccharide acceptor [2].
Family Firsts
- First stereochemistry determination
- EcOpgD by optical rotation [1].
- First general acid residue identification
- EcOpgD by X-ray crystallography and site-directed mutagenesis [1].
- First general base residue identification
- EcOpgD by X-ray crystallography and site-directed mutagenesis [1].
- First 3-D structure
- EcOpgG by X-ray crystallography [3].
References
- Motouchi S, Kobayashi K, Nakai H, and Nakajima M. (2023). Identification of enzymatic functions of osmo-regulated periplasmic glucan biosynthesis proteins from Escherichia coli reveals a novel glycoside hydrolase family. Commun Biol. 2023;6(1):961. DOI:10.1038/s42003-023-05336-6 |
- Motouchi S, Komba S, Nakai H, and Nakajima M. (2024). Discovery of Anomer-Inverting Transglycosylase: Cyclic Glucohexadecaose-Producing Enzyme from Xanthomonas, a Phytopathogen. J Am Chem Soc. 2024;146(26):17738-17746. DOI:10.1021/jacs.4c02579 |
- Hanoulle X, Rollet E, Clantin B, Landrieu I, Odberg-Ferragut C, Lippens G, Bohin JP, and Villeret V. (2004). Structural analysis of Escherichia coli OpgG, a protein required for the biosynthesis of osmoregulated periplasmic glucans. J Mol Biol. 2004;342(1):195-205. DOI:10.1016/j.jmb.2004.07.004 |