CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 52
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH52 | |
| Clan | GH-O |
| Mechanism | retaining |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/GH52.html | |
Substrate specificities
The GH52 enzymes are often isolated from various mesophilic and thermophilic bacteria, which has led to a demonstrated high thermostability within this family. The enzymes are generally monospecific, functioning as exo-β-xylosidases (EC 3.2.1.37) that cleave the terminal xylose residues from the non-reducing end of artificial xylosides and xylooligosaccharides (e.g., pNP-β-D-xylopyranoside [1, 2], xylobiose [2], and xylotriose [2]). Low levels of α-L-arabinofuranoside activity has also been observed within members of the GH52 family [3, 4], which is similar to the specificity noted for GH13 and GH54 for β-xylooligosaccharides and α-L-arabinofuranosides. The specificity for these substrates is likely due to similarities in orientation of hydroxyls and glycosidic bonds of the substrate within the active site [5, 6]. Under certain conditions, some enzymes in the family have also exhibited weak transglycosylation activity, a phenomenon that has also been infrequently observed in other glycoside hydrolases [7]. The plasticity of the active site of some GH52 members has been further explored through site-directed mutagenesis, where introduction of xylanase activity [8] and transition from a glycoside hydrolase to a glycosynthase [9] has been achieved.
Kinetics and Mechanism
Retention of stereochemistry has been observed in GH52 β-xylosidases, which is characteristic of a classical Koshland double-displacement mechanism [10]. This was first determined by Bravmen and coworkers using 1H-NMR to analyze the breakdown products of pNP-β-D-xylopyranoside by XynB2, a β-xylosidase from Bacillus stearothermophilus T-6 [1]. Further detailed analysis within this family was published in 2003 on the B. stearothermophilus XynB2 enzyme, which contained pH dependence studies (enzymatic catalysis is dependent on ionizable residues E335 and D495, with free enzyme experimental pKa values of 4.2 and 7.3, respectively) and kinetic analyses (pNP-xylobiose kcat/KM of 140 s-1mM-1; xylobiose and xylotriose KM values of 17.1x104 M-1 and 9.6x104 M-1, respectively) [4].
Catalytic Residues
Site-directed mutagenesis, chemical rescue, and kinetic profiling of XynB2 from B. stearothermophilus T-6 identified E335 as the catalytic nucleophile and D495 as the general acid/base [1, 11]. The catalytic nucleophile (E335) is conserved within the WVVNEGEY motif, which is found approximately 150 residues up-stream from the EITTYDSLD motif containing the general acid/base (D495). These results were further confirmed following the structural analysis of a GH52 from Geobacillus thermoglucosidasius [2], where in this structure the 6.5 Å separation of Glu and Asp in the active site was typical of retaining enzymes.
Three-dimensional structures
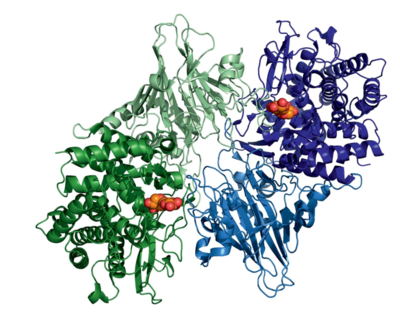
Representative structures of GH52 glycoside hydrolases have been solved, XynB2 from B. stearothermophilus T-6 (PDB ID 4RHH) and GT2_24_00240 from G. thermoglucosidasius (PDB ID 4C1P; PDB ID 4C1O). These enzymes have folds comprised of an N-terminal β-sandwich domain and a C-terminal (α/α)6 barrel domain (Figure 1) that has led to their classification into Clan GH-O, together with GH116. The exo-acting mode-of-action of GH52's is reflected in the topology of the active site. The enzymes act as dimers in solution [1, 2], with interactions between monomers of the GH52 from G. thermoglucosidasius (PDB ID 4C1P) forming a deep pocket to enclose and distort the non-reducing end xylose into a high-energy 4H3 half-chair transition conformation, while simultaneously hindering the entry of large xylan polymers into the catalytic site [2]. Furthermore, the structure of the active site also allosterically inhibits access to negative subsites beyond the -1 site. This permits interaction with only a single xylosyl residue in the negative subsites and thus hydrolysis yields a lone xylose molecule. In summary, this mechanism promotes strict exo-β-xylosidase activity, while inhibiting activity on large polymers, such as xylan.
Family Firsts
- First stereochemistry determination
- XynB2 from Bacillus stearothermophilus T-6 by 1H-NMR for the hydrolysis of pNP-β-D-xylopyranoside [1].
- First catalytic nucleophile identification
- XynB2 from Bacillus stearothermophilus T-6 by site-directed mutagenesis and chemical rescue [12].
- First general acid/base residue identification
- XynB2 from Bacillus stearothermophilus T-6 by site-directed mutagenesis, chemical rescue, and pH profiling [12].
- First 3-D structure
- GT2_24_00240 from Geobacillus thermoglucosidasius NBRC 107763 [2].
References
Error fetching PMID 24816105:
Error fetching PMID 11330658:
Error fetching PMID 12950180:
Error fetching PMID 18051350:
Error fetching PMID 1905520:
Error fetching PMID 24122394:
Error fetching PMID 25484225:
Error fetching PMID 31024890:
Error fetching PMID 12738774:
- Error fetching PMID 11322943:
- Error fetching PMID 24816105:
- Error fetching PMID 12950180:
- Error fetching PMID 18051350:
- Error fetching PMID 1905520:
- Error fetching PMID 31024890:
- Error fetching PMID 24122394:
- Error fetching PMID 25484225:
-
Koshland DE Jr: Stereochemistry and the mechanism of enzyme reactions. Biol Rev 1953, 28:416-436. DOI:10.1111/j.1469-185X.1953.tb01386.x
- Error fetching PMID 12738774:
- Error fetching PMID 11330658: