CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 93
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH93 | |
| Clan | GH-E |
| Mechanism | retaining |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/GH93.html | |
Substrate specificities
The characterized glycoside hydrolases of family GH93 are known to hydrolyse linear α-1,5-L-arabinan. [1, 2], EC:3.2.1-.
Kinetics and Mechanism
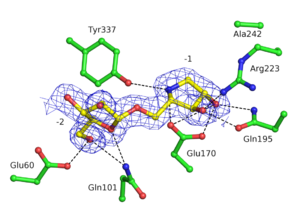
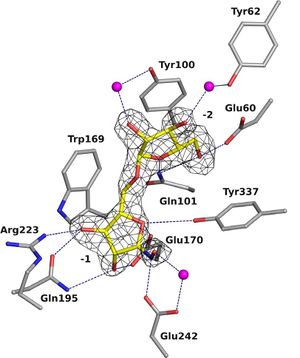
GH93 enzymes are exo-acting enzymes that only release arabinobiose from the non-reducing end of α-1,5-L-arabinan. These enzymes are proposed to be retaining enzymes based on the net retention of the configuration of the anomeric carbon is proposed from the products of the transglycosylation activity of the protein Abnx from Penicillium chrysogenum [3]. This proposal obtained support from the crystal structures of the Arb93A enzyme from Fusarium graminearum and Abnx both in complex with arabinobiose (Fig. 1) [2, 4]. α-L-Arabinofuranosylated pyrrolidines were shown to be good inhibitors of Arb93A. The Arb93A complex structure with a deoxyiminosugar equivalent of arabinobiose revealed a 4TN twist conformation expected for the Michaelis complex, as seen for several retaining GH51 α-L-arabinofuranosidases (Fig. 2) [5]. Potent shape mimic inhibitors exploiting sp2 hybridization at the anomeric carbon have been recently synthetized as well as a chromogenic substrate (Fig. 3). They are useful tools to assist further biochemical studies on L-arabinanases [6].
Catalytic Residues
From the crystal structure of Arb93A, Glu170 and Glu242 are proposed to act as catalytic nucleophile and general acid/base respectively. Mutagenesis experiment support their role in catalysis and they are strictly conserved among the family members [2]. Recent structures and mutagenesis studies for the arabinanase Abnx from Penicillium chrysogenum 31B strengthened this assignment. Mutations to alanine or glutamine of their equivalent Glu174 and Glu246 lead to inactive enzyme [4].
Three-dimensional structures
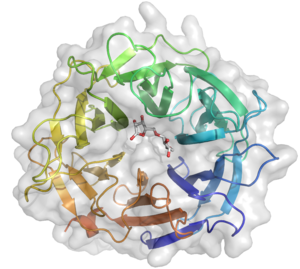
The crystal structure of Arb93A reveals a six-bladed β-propeller fold characteristic of GH33, GH34, and GH83 sialidases, which are also members of clan GH-E (Fig. 3) [2, 4]. The catalytic machinery is however very different from that of sialidases [7]. The wild-type structure was solved at 2.05 angstrom resolution in complex with arabinobiose . The active site is located in a deep acidic L-shaped crevice at the center of the beta-propeller (Fig. 1). Structures of the wild-type or E242A mutant enzyme in complex with iminoarabinobiose were solved at 1.6 and 1.85 angstrom resolution respectively as well as a complex with a shape mimic inhibitor and demonstrated ring distorsion (Fig. 2-3) [5, 6].
Family Firsts
- First sterochemistry determination
- This was determined with the Penicillium chrysogenum Abxn enzyme using 1H-NMR to identify the transglycosylation products [3]
- First catalytic nucleophile identification
- This was proposed based on the structure of Fusarium graminearum Arb93A [2]
- First general acid/base residue identification
- This was proposed based on the structure of Fusarium graminearum Arb93A [2]
- First 3-D structure
- Determined for Fusarium graminearum Arb93A by Carapito and co-workers [2]
References
Error fetching PMID 19269961:
Error fetching PMID 15342117:
Error fetching PMID 21543843:
Error fetching PMID 21773614:
Error fetching PMID 28266777:
Error fetching PMID 8591030:
- Error fetching PMID 11425761:
- Error fetching PMID 19269961:
- Error fetching PMID 15342117:
- Error fetching PMID 21543843:
- Error fetching PMID 21773614:
- Error fetching PMID 28266777:
- Error fetching PMID 8591030: