CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Polysaccharide Lyase Family 9
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Polysaccharide Lyase Family PL9 | |
| 3D Structure | β-helix |
| Mechanism | β-elimination |
| Charge neutraliser | calcium |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/PL9.html | |
Substrate specificities
Polysaccharide lyases of family 9 (CAZy) are active on pectins, a major plant cell wall polysaccharide. The main activity in characterized PL9 is pectate lyase. These enzymes cleave non-methylated α-(1-4)-linked D-galacturonic acid (homogalacturonan) by a β-elimination mechanism (EC 4.2.2.2) [1]. Two PL9 endo-acting lyases have been shown to be active on rhamnogalacturonan-I (EC 4.2.2.23) [2]. Additional activities include: exopolygalacturonic lyase (EC 4.2.2.9) and thiopeptidoglycan lyase (EC 4.2.2.-) [3, 4].
Kinetics and Mechanism
PL9 acts by an anti-β-elimination mechanism generating a 4,5-unsaturated galacturonic acid product and a new reducing end. The elimination of C5 proton is base-catalyzed by lysine 237 [1]. Similar to the PL1 family, a calcium ion interacts with the substrate carboxylate at +1 subsite promoting the C5 proton acidification. [1, 5]. The characterization of the Bacteroides thetaiotaomicron rhamnogalacturonan lyase (BT4170) revealed an additional calcium ion also interacting with the substrate and playing a role in catalysis (Figure 1) [2].
Catalytic Residues
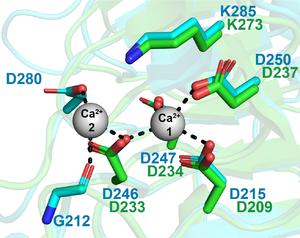
In Pel9A the lysine 237 (K237) is the Brønstead base (responsible for the abstraction of the C5 proton from galacturonic acid at +1 subsite). The calcium coordination pocket is comprised of four aspartates (D209, D233, D234 and D237) [1]. These residues are essential in catalysis and invariant in PL9 family (Figure 1) [2]. In BT4170 rhamnogalacturonan lyase, the residues G212, D246 and D280 comprise a second calcium binding site that is not conserved in pectate lyases (Figure 1) [2].
Three-dimensional structures
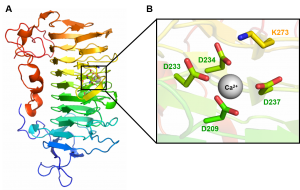
PL9 structure of Erwinia chrysanthemi (Pel9A) was solved at a resolution of 1.6 Å (PDB ID 1RU4) and displays a right-handed parallel β-helix fold (Figure 2A). The superhelical structure presents 10 complete coils and 3 β -sheets (PB1, PB2, PB3). A short α-helix at N-terminus caps the hydrophobic core of the parallel β -helix. The catalytic base K237 and calcium binding site are orientated in the structure cleft (Figure 2B) [1]. The structure of the rhamnogalacturonan lyase (BT4170) in complex with the enzyme product showed that apart from the catalytic apparatus, there is little conservation of substrate binding residues between this enzyme and the pectate lyase Pel9A [2].
Family Firsts
- First description of catalytic activity
- PelX from Erwinia chrysanthemi [3].
- First catalytic base identification
- Pel9A from Erwinia chrysanthemi [1].
- First catalytic divalent cation identification
- Pel9A from Erwinia chrysanthemi [1].
- First 3-D structure
- Pel9A from Erwinia chrysanthemi [1].
References
- Jenkins J, Shevchik VE, Hugouvieux-Cotte-Pattat N, and Pickersgill RW. (2004). The crystal structure of pectate lyase Pel9A from Erwinia chrysanthemi. J Biol Chem. 2004;279(10):9139-45. DOI:10.1074/jbc.M311390200 |
- Luis AS, Briggs J, Zhang X, Farnell B, Ndeh D, Labourel A, Baslé A, Cartmell A, Terrapon N, Stott K, Lowe EC, McLean R, Shearer K, Schückel J, Venditto I, Ralet MC, Henrissat B, Martens EC, Mosimann SC, Abbott DW, and Gilbert HJ. (2018). Dietary pectic glycans are degraded by coordinated enzyme pathways in human colonic Bacteroides. Nat Microbiol. 2018;3(2):210-219. DOI:10.1038/s41564-017-0079-1 |
- Brooks AD, He SY, Gold S, Keen NT, Collmer A, and Hutcheson SW. (1990). Molecular cloning of the structural gene for exopolygalacturonate lyase from Erwinia chrysanthemi EC16 and characterization of the enzyme product. J Bacteriol. 1990;172(12):6950-8. DOI:10.1128/jb.172.12.6950-6958.1990 |
- Kondo K, Takeda M, Ejima W, Kawasaki Y, Umezu T, Yamada M, Koizumi J, Mashima T, and Katahira M. (2011). Study of a novel glycoconjugate, thiopeptidoglycan, and a novel polysaccharide lyase, thiopeptidoglycan lyase. Int J Biol Macromol. 2011;48(2):256-62. DOI:10.1016/j.ijbiomac.2010.11.009 |
- Seyedarabi A, To TT, Ali S, Hussain S, Fries M, Madsen R, Clausen MH, Teixteira S, Brocklehurst K, and Pickersgill RW. (2010). Structural insights into substrate specificity and the anti beta-elimination mechanism of pectate lyase. Biochemistry. 2010;49(3):539-46. DOI:10.1021/bi901503g |