CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Polysaccharide Lyase Family 22
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Polysaccharide Lyase Family PL22 | |
| 3D Structure | β7 propeller |
| Mechanism | β-elimination |
| Charge neutraliser | manganese |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/PL22.html | |
Substrate specificities
Family 22 Polysaccharide Lyases (PL22s) contain two subfamilies and several outlier sequences [1]. Originally known as oligogalacturonide transeliminases (OGTE) [2], PL22s are now commonly referred to as oligogalacturonide lyases (OGLs). This enzyme family is found primarily in phytopathogenic or intestinal bacteria where it plays a role in the metabolism of pectin.
PL22s remove 5-keto-4-deoxyuronate (4-deoxy-l-threo-5-hexosulose uronic acid, DKI) from short chain oligalacturonides and display preferential activity on digalacturonate and Δ4,5-unsaturated digalacturonate [3, 4]. Activity on trigalacturonate is significantly lower and PL22s appear to completely lack activity on long chain polymers of α-(1,4)-linked polygalacturonate [3, 4]. Differing levels of activity has been reported on methylated short chain oligogalacturonides depending on the location of methylation [4].
Kinetics and Mechanism
PL22s harness a β-elimination mechanism to cleave the glycosidic bonds in oligogalacturonides. This process requires a Brønstead base for proton abstraction and a catalytic metal (e.g. Mn2+ or Mg2+) for acidification of the α-proton and charge neutralization of the oxyanion intermediate. YePL22 (YE1876 from Yersinia enterocolitica subsp. enterocolitica 8081; gi|123442156|) displays the lowest reported pH optimum for a pectate lyase (7.3 - 7.7) [3], which is substantially lower than other families that deploy catalytic arginines or lysines in the β-elimination of pectate.
Catalytic Residues
Within the structure of YePL22, H242 is the only basic residue that is in proximity to the α-proton of a modelled galacturonate [3]. This histidine is highly conserved within PL22s with only Candidatus Solibacter usitatus Ellin6076 (gi|116225114|) displaying an alternative residue (T236); however, whether this protein is a lyase has yet to be determined. The 'stabilizing arginine' [3] (YE1876: R217) is completely conserved across the PL22 family.
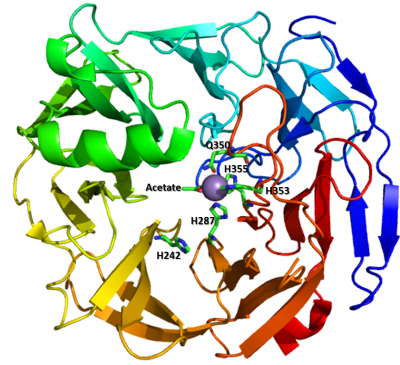
The metal coordination pocket houses a manganese ion and is comprised of three histidines (VPA0088: H287, H353, H355; YE1876: H287, H353, H355) and one glutamine (VPA0088: Q350; YeOGL: Q350). It is of note however that although these residues are perfectly conserved in all reported subfamily 1 sequences and several outlier sequences, there are minor differences in subfamily 2 [1]: H287 is invariant, Q350 is not conserved, and H353 and H355 have been replaced with a glutamate and asparagine respectively. These modifications may alter the chemistry of metal coordination selectivity. Further experimentation will be required to define this relationship.
Three-dimensional structures
The first structure of a PL22 determined was from Vibrio parahaemolyticus RIMD 2210633 (PDB 3C5M) in 2008 by x-ray diffraction to 2.60 Å (Northeast Structural Genomics Consortium). This was followed in 2010 by YePL2A from Yersinia enterocolitica subsp. enterocolitica 8081 (PDB 3PE7), which was solved in complex with Mn2+ and acetate by x-ray diffraction to 1.65 Å [3]. The two proteins share ~69% sequence identity and highly similar 3D structures. The PL22 fold is a β7 propeller with the catalytic machinery and metal coordination pocket housed at the center of the enzyme.
Family Firsts
- First catalytic activity
- OGTE from Pectobacterium carotovorum ICPB EC153 (previously Erwinia carotovora) [2].
- First catalytic base identification
- YeOGL (YE1876) H242 from Yersinia enterocolitica subsp. enterocolitica 8081 [3].
- First catalytic divalent cation identification
- OGL (Dda3937_03686) from Dickeya Dadantii 3937 (previously Erwinia chrysanthemi 3937) [5].
- First 3-D structure
- VPA0088 from Vibrio parahaemolyticus RIMD 2210633 (Unpublished: PDB 3C5M).
References
- Lombard V, Bernard T, Rancurel C, Brumer H, Coutinho PM, and Henrissat B. (2010). A hierarchical classification of polysaccharide lyases for glycogenomics. Biochem J. 2010;432(3):437-44. DOI:10.1042/BJ20101185 |
- Moran F, Nasuno S, and Starr MP. (1968). Oligogalacturonide trans-eliminase of Erwinia carotovora. Arch Biochem Biophys. 1968;125(3):734-41. DOI:10.1016/0003-9861(68)90508-0 |
- Abbott DW, Gilbert HJ, and Boraston AB. (2010). The active site of oligogalacturonate lyase provides unique insights into cytoplasmic oligogalacturonate beta-elimination. J Biol Chem. 2010;285(50):39029-38. DOI:10.1074/jbc.M110.153981 |
- Kester HC, Magaud D, Roy C, Anker D, Doutheau A, Shevchik V, Hugouvieux-Cotte-Pattat N, Benen JA, and Visser J. (1999). Performance of selected microbial pectinases on synthetic monomethyl-esterified di- and trigalacturonates. J Biol Chem. 1999;274(52):37053-9. DOI:10.1074/jbc.274.52.37053 |
- Shevchik VE, Condemine G, Robert-Baudouy J, and Hugouvieux-Cotte-Pattat N. (1999). The exopolygalacturonate lyase PelW and the oligogalacturonate lyase Ogl, two cytoplasmic enzymes of pectin catabolism in Erwinia chrysanthemi 3937. J Bacteriol. 1999;181(13):3912-9. DOI:10.1128/JB.181.13.3912-3919.1999 |
- Collmer A and Bateman DF. (1981). Impaired induction and self-catabolite repression of extracellular pectate lyase in Erwinia chrysanthemi mutants deficient in oligogalacturonide lyase. Proc Natl Acad Sci U S A. 1981;78(6):3920-4. DOI:10.1073/pnas.78.6.3920 |
- Reverchon S and Robert-Baudouy J. (1987). Molecular cloning of an Erwinia chrysanthemi oligogalacturonate lyase gene involved in pectin degradation. Gene. 1987;55(1):125-33. DOI:10.1016/0378-1119(87)90255-1 |
- Reverchon S, Huang Y, Bourson C, and Robert-Baudouy J. (1989). Nucleotide sequences of the Erwinia chrysanthemi ogl and pelE genes negatively regulated by the kdgR gene product. Gene. 1989;85(1):125-34. DOI:10.1016/0378-1119(89)90472-1 |
- Yang S, Zhang Q, Guo J, Charkowski AO, Glick BR, Ibekwe AM, Cooksey DA, and Yang CH. (2007). Global effect of indole-3-acetic acid biosynthesis on multiple virulence factors of Erwinia chrysanthemi 3937. Appl Environ Microbiol. 2007;73(4):1079-88. DOI:10.1128/AEM.01770-06 |