CAZypedia needs your help! We have many unassigned GH, PL, CE, AA, GT, and CBM pages in need of Authors and Responsible Curators.
Scientists at all career stages, including students, are welcome to contribute to CAZypedia. Read more here, and in the 10th anniversary article in Glycobiology.
New to the CAZy classification? Read this first.
*
Consider attending the 15th Carbohydrate Bioengineering Meeting in Ghent, 5-8 May 2024.
Difference between revisions of "Carbohydrate Binding Module Family 89"
| (18 intermediate revisions by 4 users not shown) | |||
| Line 1: | Line 1: | ||
<!-- RESPONSIBLE CURATORS: Please replace the {{UnderConstruction}} tag below with {{CuratorApproved}} when the page is ready for wider public consumption --> | <!-- RESPONSIBLE CURATORS: Please replace the {{UnderConstruction}} tag below with {{CuratorApproved}} when the page is ready for wider public consumption --> | ||
| − | {{ | + | {{CuratorApproved}} |
* [[Author]]s: [[User:Mariana Morais|Mariana Abrahão Bueno de Morais]] and [[User:Gabriela Persinoti|Gabriela Felix Persinoti]] | * [[Author]]s: [[User:Mariana Morais|Mariana Abrahão Bueno de Morais]] and [[User:Gabriela Persinoti|Gabriela Felix Persinoti]] | ||
* [[Responsible Curator]]: [[User:Mario Murakami|Mario Murakami]] | * [[Responsible Curator]]: [[User:Mario Murakami|Mario Murakami]] | ||
| Line 17: | Line 17: | ||
== Ligand specificities == | == Ligand specificities == | ||
| − | CBM89 | + | The CBM89 family was created based on the discovery of an N-terminal domain appended to a [[GH10]] xylanase recovered from a metagenome-assembled genome (MAG) of the capybara gut microbiota. The founding member, CapCBM89, was shown to interact with beechwood xylan and rye arabinoxylan according to affinity gel electrophoresis (AGE) <cite>Cabral2022</cite>. Although similar β-helix structural architectures have been found in the families [[GH28]], [[GH91]], [[PL6]], and [[CE8]], no catalytic activity of the isolated CapCBM89 was observed on the substrates tested <cite>Cabral2022</cite>. |
== Structural Features == | == Structural Features == | ||
| − | + | [[File:Cbm89_cazypedia.png|thumb|300px|right|'''Figure 1.''' CapCBM89 structure (PDB 7JVI [https://www.rcsb.org/structure/7JVI]). The Ca<sup>2+</sup> (orange sphere), its coordinating residue (D344) and the two residues that impaired polysaccharide binding (Y62 and E82) are highlighted.]] | |
| − | + | The crystal structure of the founding member was solved at 1.85 Å resolution, which consists of 14 helical turns establishing a parallel β-helix fold <cite>Cabral2022</cite>. A Ca<sup>2+</sup> was found in the structure, connecting the 11<sup>th</sup> and 12<sup>th</sup> helical turns ([{{PDBlink}}7jvi PDB 7JVI]). The role of the ion seems to be only related to the domain stabilization, since a mutation in the Ca<sup>2+</sup> binding site (D344L) affected the protein stability but not its capacity to bind to arabinoxylan/xylan <cite>Cabral2022</cite>. In CapCBM89, mutants Y62A and E82A impaired the capacity to bind to arabinoxylan/xylan, indicating the importance of this motif for carbohydrate binding <cite>Cabral2022</cite> . | |
| − | |||
== Functionalities == | == Functionalities == | ||
| − | + | In general, CBM89 members comprise approximately 600 to 1000 residues, which can be fused to GH domains. The most common associated glycoside hydrolases are from the [[GH10]] family. The founding member, [https://www.ncbi.nlm.nih.gov/protein/UMW87592.1 CapCBM89], was found appended to the N-terminus of a [[GH10]] domain, which is clustered with CAZymes targeting arabinoxylans <cite>Cabral2022</cite>. This CAZyme cluster encodes the CapCBM89-[[GH10]] and two exo-acting enzymes from [https://www.ncbi.nlm.nih.gov/protein/2205462650 GH43] and [https://www.ncbi.nlm.nih.gov/protein/2205462646 GH97] families <cite>Cabral2022</cite>. The [[GH10]] domain is active on arabinoxylan and xylan, which is in agreement with the binding assays of CapCBM89, suggesting that CapCBM89 targets the [[GH10]] domain to these substrates for catalysis <cite>Cabral2022</cite>. | |
| − | |||
| + | == Family Firsts == | ||
| + | ;First Identified: The first identified CBM89 member (CapCBM89) was from an uncultured Bacteroidaceae MAG57 recovered from the capybara gut microbiome <cite>Cabral2022</cite> . | ||
| − | + | ;First Structural Characterization: The first CBM89 structure is [{{PDBlink}}7jvi PDB 7JVI], retrieved from the capybara gut microbiome <cite>Cabral2022</cite> . | |
| − | |||
| − | |||
| − | |||
| − | The first CBM89 structure is | ||
== References == | == References == | ||
<biblio> | <biblio> | ||
| − | #Cabral2022 pmid= 35110564 | + | #Cabral2022 pmid=35110564 |
</biblio> | </biblio> | ||
Latest revision as of 03:21, 6 July 2023
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
- Authors: Mariana Abrahão Bueno de Morais and Gabriela Felix Persinoti
- Responsible Curator: Mario Murakami
| CAZy DB link | |
| http://www.cazy.org/CBM89.html |
Ligand specificities
The CBM89 family was created based on the discovery of an N-terminal domain appended to a GH10 xylanase recovered from a metagenome-assembled genome (MAG) of the capybara gut microbiota. The founding member, CapCBM89, was shown to interact with beechwood xylan and rye arabinoxylan according to affinity gel electrophoresis (AGE) [1]. Although similar β-helix structural architectures have been found in the families GH28, GH91, PL6, and CE8, no catalytic activity of the isolated CapCBM89 was observed on the substrates tested [1].
Structural Features
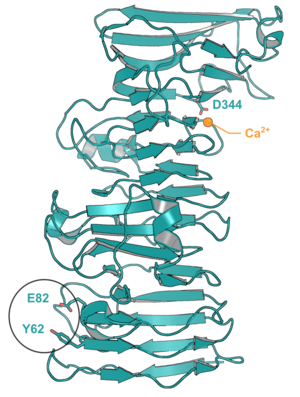
The crystal structure of the founding member was solved at 1.85 Å resolution, which consists of 14 helical turns establishing a parallel β-helix fold [1]. A Ca2+ was found in the structure, connecting the 11th and 12th helical turns (PDB 7JVI). The role of the ion seems to be only related to the domain stabilization, since a mutation in the Ca2+ binding site (D344L) affected the protein stability but not its capacity to bind to arabinoxylan/xylan [1]. In CapCBM89, mutants Y62A and E82A impaired the capacity to bind to arabinoxylan/xylan, indicating the importance of this motif for carbohydrate binding [1] .
Functionalities
In general, CBM89 members comprise approximately 600 to 1000 residues, which can be fused to GH domains. The most common associated glycoside hydrolases are from the GH10 family. The founding member, CapCBM89, was found appended to the N-terminus of a GH10 domain, which is clustered with CAZymes targeting arabinoxylans [1]. This CAZyme cluster encodes the CapCBM89-GH10 and two exo-acting enzymes from GH43 and GH97 families [1]. The GH10 domain is active on arabinoxylan and xylan, which is in agreement with the binding assays of CapCBM89, suggesting that CapCBM89 targets the GH10 domain to these substrates for catalysis [1].
Family Firsts
- First Identified
- The first identified CBM89 member (CapCBM89) was from an uncultured Bacteroidaceae MAG57 recovered from the capybara gut microbiome [1] .
- First Structural Characterization
- The first CBM89 structure is PDB 7JVI, retrieved from the capybara gut microbiome [1] .
References
- Cabral L, Persinoti GF, Paixão DAA, Martins MP, Morais MAB, Chinaglia M, Domingues MN, Sforca ML, Pirolla RAS, Generoso WC, Santos CA, Maciel LF, Terrapon N, Lombard V, Henrissat B, and Murakami MT. (2022). Gut microbiome of the largest living rodent harbors unprecedented enzymatic systems to degrade plant polysaccharides. Nat Commun. 2022;13(1):629. DOI:10.1038/s41467-022-28310-y |