CAZypedia needs your help! We have many unassigned GH, PL, CE, AA, GT, and CBM pages in need of Authors and Responsible Curators.
Scientists at all career stages, including students, are welcome to contribute to CAZypedia. Read more here, and in the 10th anniversary article in Glycobiology.
New to the CAZy classification? Read this first.
*
Consider attending the 15th Carbohydrate Bioengineering Meeting in Ghent, 5-8 May 2024.
Polysaccharide Lyase Family 9
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Ana Luis^^^
- Responsible Curator: ^^^Wade Abbott^^^
| Polysaccharide Lyase Family PL9 | |
| 3D Structure | β-helix |
| Mechanism | β-elimination |
| Charge neutraliser | calcium |
| Active site residues | known |
| CAZy DB link | |
| http://www.cazy.org/PL9.html | |
Substrate specificities
Polysaccharide lyases of family 9 (CAZy) degrade homogalacturonan,a pectin component present in the plant cell walls. An enzyme in PL9 was described as active on sheath, a thioloic glycoconjugate secreted by Sphaerotilus natans [1]. The main activity in characterized PL9 is pectate lyases. These enzymes cleave non-methylated α-(1-4)-linked D-galacturonic acid by a β-elimination mechanism (EC 4.2.2.2) [2]. Additional activities include: exopolygalacturonic lyase (EC 4.2.2.9) and thiopeptidoglycan lyase (EC 4.2.2.-) [3, 4].
Kinetics and Mechanism
PL9 acts by an anti-β-elimination mechanism generating a 4,5-unsaturated galacturonic acid product and a new reducing end. The elimination of C5 proton is base-catalyzed by lysine 237 [2]. Similar to the PL1 family, a calcium ion interacts with the substrate carboxylate at +1 subsite promoting the C5 proton acidification. [2, 5].
Catalytic Residues
The lysine 237 (K237) is the Brønstead base (responsible for the abstraction of the C5 proton from galacturonic acid at +1 subsite). The calcium coordination pocket is comprised of four aspartates (D209, D233, D234 and D237) [2].
Three-dimensional structures
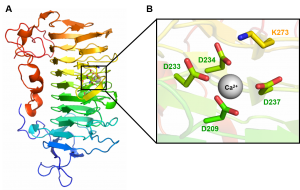
PL9 structure of Erwinia chrysanthemi (Pel9A) was solved at a resolution of 1.6 Å (PDB ID 1RU4) and displays a right-handed parallel β-helix fold (Figure 1A). The superhelical structure presents 10 complete coils and 3 β -sheets (PB1, PB2, PB3). A short α-helix at N-terminus caps the hydrophobic core of the parallel β -helix. The catalytic base K237 and calcium binding site are orientated in the structure cleft (Figure 1B) [2].
Family Firsts
- First description of catalytic activity
- PelX from Erwinia chrysanthemi [3].
- First catalytic base identification
- Pel9A from Erwinia chrysanthemi [2].
- First catalytic divalent cation identification
- Pel9A from Erwinia chrysanthemi [2].
- First 3-D structure
- Pel9A from Erwinia chrysanthemi [2].
References
Error fetching PMID 11055955:
Error fetching PMID 21095202:
Error fetching PMID 2254266:
- Error fetching PMID 11055955:
- Error fetching PMID 14670977:
- Error fetching PMID 2254266:
- Error fetching PMID 21095202:
- Seyedarabi A, To TT, Ali S, Hussain S, Fries M, Madsen R, Clausen MH, Teixteira S, Brocklehurst K, and Pickersgill RW. (2010). Structural insights into substrate specificity and the anti beta-elimination mechanism of pectate lyase. Biochemistry. 2010;49(3):539-46. DOI:10.1021/bi901503g |