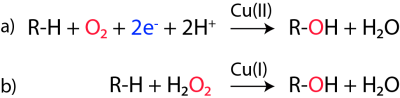CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Auxiliary Activity Family 10
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Auxiliary Activity Family 10 | |
| Clan | Structurally related to AA9 & AA11 & AA13 |
| Mechanism | lytic oxidase |
| Active site residues | mononuclear copper ion |
| CAZy DB link | |
| https://www.cazy.org/AA10.html | |
Substrate specificities
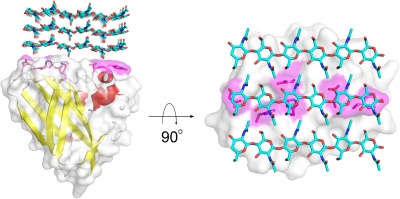
Members of the AA10 family of lytic polysaccharide monooxygenases were previously classified as Carbohydrate Binding Module Family 33 members). They are known to cleave chitin [3, 4, 5] and cellulose [6, 7]. Most characterized AA10 LPMO oxidize the C1 of the scissile glycosidic bond, but some also oxidize C4 [7, 8]. AA10 LPMOs are closely related to LPMOs in families AA11 and AA13, known to cleave chitin and starch, respectively. AA10 modules often occur in combination with additional modules, in particular carbohydrate-binding modules (CBMs), but also catalytic domains such as GH18 chitinases [2]. The CBMs contribute to substrate-binding and may also affect operational stability of the LPMO [8, 9, 10].
Before proteins belonging to AA10 were identified as enzymes, they were generally known as chitin binding proteins (CBPs) and as carbohydrate-binding modules belonging to family CBM33. The reason for this was that most AA10s studied had been identified in chitinolytic systems such as that of Serratia marcescens [11, 12], several Streptomyces species [13, 14, 15], Bacillus amyloliquefaciens [16],Vibrio cholerae [17], Pseudomonas aeruginosa [18] and Lacotococcus lactis [19]. Upon their characterization no other function than substrate binding could be identified, thus the name "chitin binding protein" was coined. Substrates and potential substrates identified by binding studies include alpha-chitin [13, 14], beta-chitin [1, 12], both the alpha- and beta-chitin allomorphs [4, 16, 19] chitosan [15], cellulose [6, 20] and even bacterial and eptihelial cell surfaces where the binding interaction substrate has been suggested to be GlcNAc-containing glycoproteins or proteoglycans [17, 21]. It should be noted that studies on AA10s prior to their identification as copper-dependent metalloenzymes were conducted in the absence of Cu(II), which may have had influence on the binding affinity and specificity of the enzyme.
As previously noted, AA10s exist both as single module entities and in multimodular forms. CBP21 from S. marcescens [3] and EfCBM33A from E. faecalis [4] are catalytically functional single-module AA10s that both bind well to chitin (CBP21 is specific for beta-chitin, whereas EfCBM33A binds to both alpha and beta-chitin). The cellulose targeting AA10 from S. coelicolor (CelS2) on the other hand, is bimodular and contains a cellulose binding CBM2 in addition to the catalytic AA10 module [6, 7, 9]. The V. cholerae AA10 is an elongated tetra-modular protein where the N-terminal catalytically active AA10 module binds mucin, the two following modules bind bacterial cell walls and the C-terminal CBM5/12 binds chitin [17, 22].
The substrate binding surface of AA10s is flat and lacks the typical arrangement of aromatic amino acids that is common for carbohydrate binding proteins [1, 23]. It is therefore thought that substrate binding is predominantly mediated by hydrogen bonds, as is substantiated by a recent study of CelS2 [8]. The flat substrate binding surface seems optimal for binding the flat surface of crystalline carbohydrate structures like cellulose and chitin (Fig. 1).
Kinetics and Mechanism
The catalytic mechanism of LPMOs is a matter of intense research and debate. Using labeled water and molecular oxygen, Vaaje-Kolstad et al [3] showed that one LPMO reaction requires two externally delivered electrons and molecular oxygen (Figure 2). Phillips et al [24] pointed out that hydrogen abstraction (from the C1 or the C4) by a reactive oxygen species coordinated by the copper would be followed by hydroxylation and that this would lead to spontaneous cleavage of the glycosidic bond, leaving either the C1 or the C4 oxidized (yielding a lactone in equilibrium with carboxylic acid or a 4-keto group in equilibrium with a gemdiol, respectively [25]). The nature of the hydrogen abstracting oxygen species is not known. Current literature seems to be converging towards a Cu(II)-oxyl species [24, 26, 27, 28, 29], but there are plausible alternatives that cannot be excluded, as outlined in e.g. [27, 29, 30, 31].
One challenging element of understanding how LPMOs work relates to the required delivery of two electrons to a substrate-bound enzyme whose single co-factor only allows storage of one electron. While several scenarios have been proposed, a radical and experimentally supported solution to this problem was recently proposed by Bissaro et al [27]. As shown in Fig. 2, panel b, the proposal is that hydrogen peroxide rather than molecular oxygen is the preferred co-substrate of LPMOs. The H2O2-based mechanism would only require a priming reduction of the LPMO to its Cu(I) state, which would then be capable of carrying out multiple catalytic cycles while having its electrons and oxygen delivered in the form of partially reduced O2, namely H2O2. Bissaro et al. point out that H2O2 will be generated under the reaction conditions that are normally used to assess LPMO functionality. Further work is needed to assess the validity of the various proposed LPMO mechanisms. One key challenge in such work is that LPMOs become rapidly inactivated due to oxidative damage in the catalytic center [27, 32]
Reported catalytic rates of LPMOs, including AA10 LPMOs, under standard conditions (= the presence of a reductant and O2), are remarkably low and tend to vary from 1 to 10 per minute (e.g. [3, 33, 34]). Although not usually reported in the literature, progress curves for LPMO reactions are notoriously non-linear and there is thus little available kinetic data (see [32, 34] for relatively high quality kinetic data). Importantly, it has been shown that LPMO rates can be drastically enhanced by using light [35] or H2O2 [27] to drive the LPMO reaction. In a recent detailed kinetic study of CBP21, using hydrogen peroxide to drive the reaction, Kuusk et al [32] determined the kcat for chitin oxidation to be 5.6 s-1, and concluded that the kcat/Km for H2O2-driven degradation of chitin was in the order of 106 M-1 s-1.
Catalytic Residues and copper coordination
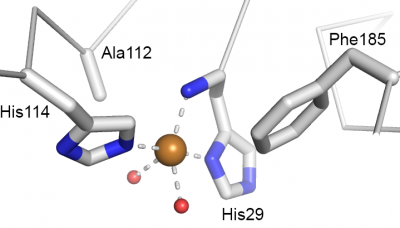
Catalysis happens at the single catalytic copper ion that is coordinated by a so-called histidine brace [37] (Fig. 3) that is strictly conserved in all LPMOs. Originally, there was some confusion about the nature of the metal ion [3, 38], which is due to the high copper affinity of LPMOs, which allows the enzymes to acquire copper even in the "absence" of copper. This issue was settled for AA9 LPMOs by Quinlan et al [37] and Phillips et al [24] and further elaborated for AA10s in subsequent publications [5, 39]. The two histidines making up the histidine brace coordinate the copper in three positions that, for practical reasons are often referred to as "equatorial", although this is not always formally correct (as exemplified by Fig. 3). One of the histidines is the N-terminus of the mature protein and contributes with both its N-terminal amino group and a its imidazole side chain, whereas the other histidine contribute only with its side chain. In oxidized LPMOs, i.e. the Cu(II) form, the fourth equatorial position is occupied by a water [36, 40]. There are strong indications that the reactive oxygen species formed during catalysis emerges at this fourth equatorial position [34, 41]. The final two coordination positions, often referred to as axial, vary. The proximal axial coordination site (Fig. 3) is occupied by a phenylalanine or a tyrosine. The functional implications, if any, of this difference remains unknown. It should be noted that, while the hydroxyl group of the tyrosine does point towards the copper ion, the distance to the copper is too long for a strong interaction. The distal, solvent-exposed axial position tends to be occupied by a water in oxidized LPMOs and recent studies on AA9s [34, 42] suggest that this water will be displaced by substrate binding. The copper ion is partly shielded from the solvent in this axial position by a conserved alanine residue, the role of which has received considerable attention in the literature [7, 8, 40]. Current data do not point at a particular effect of the presence of this alanine nor an effect of its exact position, which varies among AA10 LPMOs.
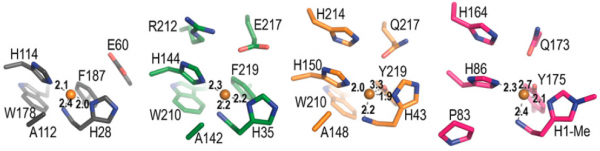
The immediate environment of the copper binding site, sometimes referred to as the second shell, shows functionally conserved features that are known from experiment to be important for catalysis (e.g. [8, 43, 44]). Further analysis and discussion of the roles of these residues awaits deeper insights into the catalytic mechanism of LPMOs and better assays for testing the true catalytic potential of LPMO variants. Figure 4 shows an example of such functionally conserved additional structural features and reference [8] describes a recent mutagenesis study on an AA10, including a discussion of the many pitfalls when trying to functionally interprete mutational effects.
Three-dimensional structures
In 2005 the structure CBP21 from S. marcescens was solved and represents the first structure in the AA10 family 2BEM [1] and the first structure of an LPMO. The CBP21 wild type structure has three molecules in the asymetric unit, of which only chain C shows electron density for a metal bound in the metal binding motif (modeled as a sodium ion, but, in retrospect this is probably a reduced copper ion with low occupancy). Later the same year the structure of the CBP21-Y54A mutant was solved (different crystal form and space group), showing two molecules in the asymetric unit with no trace of electron density for a metal ion bound in the active site 2BEN [43]. The second AA10 structure, one of two AA10 from Burkholderia pseudomallei 1710b (Uniprot ID: Q3JY22), was published in the PDB late in 2011 by the Seattle Structural Genomics Center for Infectious Disease 3UAM. The structure contains five molecules in the asymetric unit that all have two amino acids from the signal peptide still attached to the N-terminus, most likely disrupting the active site. The third unique AA10 structure to be solved was GbpA from Vibrio cholerae O1 biovar El Tor str. N16961 2XWX [17]. GbpA is unique in the sense that it contains four discrete modules: a catalytically active N-terminal AA10 module, two modules likely involved in binding to bacterial cell walls and a C-terminal CBM5/12. The published structure of GbpA only lacks the C-termainal CBM5/12 and is thus the first multimodular AA10 structure to be published. Shortly after the release of the GbpA, the structure of EfCBM33A from Enterococcus faecalis 4A02 [4] was published. The structure of EfCBM33A was solved at very high resolution (0.95Å), but similar to the other structures solved, no metal ion was observed bound in the active site. In 2012 the solution structure of CBP21 wild type (apo-form) was solved by NMR 2LHS [5], and this study also provided evidence for copper being the active site metal, as had been previously shown for AA11 LPMOs [37]. In spring 2013 the structure of the single AA10 harbored byBacillus amyloliquefaciens DSM7 (BaCBM33) was published [39]. The latter publication contained three structures: the apo-enzyme 2YOW and two structures containing a reduced copper ion bound to the active site 2YOY2YOX, thus being the first AA10 structures with a copper ion bound in the active site. The first structures of AA10 LPMOs known to be active on cellulose came in 2014 [7]. It is worth noting that the oxidation state of the copper ion is affected by the reducing power of the X-ray beam as discussed by Hemsworth et al [40] and Gudmundsson et al [36].
Family Firsts
- First AA10 protein identified
- The first proteins studied from the AA10 family were all isolated and cloned from various Streptomyces strains, a major effort carried out by the Schrempf group of Osnabrück University. The first family AA10 protein to be isolated and characterized was CHB1 from Streptomyces olivaceoviridis a study published in 1994 [45]. CHB1 was shown to bind strongly to alpha-chitin and was also observed to bind to fungal hyphae. When these proteins originally were included into CAZy, they were classified as CBM33.
- First demonstration of synergy between AA10 and canonical glycoside hydrolases
- In 2005, Vaaje-Kolstad and co-workers showed that CBP21 from S. marcescens is able to increase the rate of chitin hydrolysis by a variety of chitinases, including all three S. marsescens GH18 chitinases and a GH19 chitinase from Streptomyces coelicolor [43]. It is worth noting that the title of the original publication erroneously qualified CBP21 as "non-catalytic"; this is due to how one was thinking about possible substrate-disrupting effects of CBMs at the time.
- First demonstration of oxidative cleavage by an AA10 protein
- Catalysis of lytic oxidation of a glycosidic bond by an AA10 enzyme was first shown for CBP21 in 2010, where oxidative cleavage of chitin was demonstrated [3]. This finding also represents the first demonstration of the true nature of LPMO activity, regardless of AA family. Oxidative cleavage of cellulose by an AA10 was demonstrated by the same group in 2011 [6].
- First 3-D structure
- CBP21, the single AA10-type LPMO from the Gram negative bacterium Serratia marcescens, PDB ID 2BEM.
References
- Vaaje-Kolstad G, Houston DR, Riemen AH, Eijsink VG, and van Aalten DM. (2005). Crystal structure and binding properties of the Serratia marcescens chitin-binding protein CBP21. J Biol Chem. 2005;280(12):11313-9. DOI:10.1074/jbc.M407175200 |
- Horn SJ, Vaaje-Kolstad G, Westereng B, and Eijsink VG. (2012). Novel enzymes for the degradation of cellulose. Biotechnol Biofuels. 2012;5(1):45. DOI:10.1186/1754-6834-5-45 |
- Vaaje-Kolstad G, Westereng B, Horn SJ, Liu Z, Zhai H, Sørlie M, and Eijsink VG. (2010). An oxidative enzyme boosting the enzymatic conversion of recalcitrant polysaccharides. Science. 2010;330(6001):219-22. DOI:10.1126/science.1192231 |
- Vaaje-Kolstad G, Bøhle LA, Gåseidnes S, Dalhus B, Bjørås M, Mathiesen G, and Eijsink VG. (2012). Characterization of the chitinolytic machinery of Enterococcus faecalis V583 and high-resolution structure of its oxidative CBM33 enzyme. J Mol Biol. 2012;416(2):239-54. DOI:10.1016/j.jmb.2011.12.033 |
- Aachmann FL, Sørlie M, Skjåk-Bræk G, Eijsink VG, and Vaaje-Kolstad G. (2012). NMR structure of a lytic polysaccharide monooxygenase provides insight into copper binding, protein dynamics, and substrate interactions. Proc Natl Acad Sci U S A. 2012;109(46):18779-84. DOI:10.1073/pnas.1208822109 |
- Forsberg Z, Vaaje-Kolstad G, Westereng B, Bunæs AC, Stenstrøm Y, MacKenzie A, Sørlie M, Horn SJ, and Eijsink VG. (2011). Cleavage of cellulose by a CBM33 protein. Protein Sci. 2011;20(9):1479-83. DOI:10.1002/pro.689 |
- Forsberg Z, Mackenzie AK, Sørlie M, Røhr ÅK, Helland R, Arvai AS, Vaaje-Kolstad G, and Eijsink VG. (2014). Structural and functional characterization of a conserved pair of bacterial cellulose-oxidizing lytic polysaccharide monooxygenases. Proc Natl Acad Sci U S A. 2014;111(23):8446-51. DOI:10.1073/pnas.1402771111 |
- Forsberg Z, Bissaro B, Gullesen J, Dalhus B, Vaaje-Kolstad G, and Eijsink VGH. (2018). Structural determinants of bacterial lytic polysaccharide monooxygenase functionality. J Biol Chem. 2018;293(4):1397-1412. DOI:10.1074/jbc.M117.817130 |
- Forsberg Z, Røhr AK, Mekasha S, Andersson KK, Eijsink VG, Vaaje-Kolstad G, and Sørlie M. (2014). Comparative study of two chitin-active and two cellulose-active AA10-type lytic polysaccharide monooxygenases. Biochemistry. 2014;53(10):1647-56. DOI:10.1021/bi5000433 |
- Crouch LI, Labourel A, Walton PH, Davies GJ, and Gilbert HJ. (2016). The Contribution of Non-catalytic Carbohydrate Binding Modules to the Activity of Lytic Polysaccharide Monooxygenases. J Biol Chem. 2016;291(14):7439-49. DOI:10.1074/jbc.M115.702365 |
- Fuchs RL, McPherson SA, and Drahos DJ. (1986). Cloning of a Serratia marcescens Gene Encoding Chitinase. Appl Environ Microbiol. 1986;51(3):504-9. DOI:10.1128/aem.51.3.504-509.1986 |
- Suzuki K, Suzuki M, Taiyoji M, Nikaidou N, and Watanabe T. (1998). Chitin binding protein (CBP21) in the culture supernatant of Serratia marcescens 2170. Biosci Biotechnol Biochem. 1998;62(1):128-35. DOI:10.1271/bbb.62.128 |
- Zeltins A and Schrempf H. (1997). Specific interaction of the Streptomyces chitin-binding protein CHB1 with alpha-chitin--the role of individual tryptophan residues. Eur J Biochem. 1997;246(2):557-64. DOI:10.1111/j.1432-1033.1997.t01-1-00557.x |
- Kolbe S, Fischer S, Becirevic A, Hinz P, and Schrempf H. (1998). The Streptomyces reticuli alpha-chitin-binding protein CHB2 and its gene. Microbiology (Reading). 1998;144 ( Pt 5):1291-1297. DOI:10.1099/00221287-144-5-1291 |
- Saito A, Miyashita K, Biukovic G, and Schrempf H. (2001). Characteristics of a Streptomyces coelicolor A3(2) extracellular protein targeting chitin and chitosan. Appl Environ Microbiol. 2001;67(3):1268-73. DOI:10.1128/AEM.67.3.1268-1273.2001 |
- Chu HH, Hoang V, Hofemeister J, and Schrempf H. (2001). A Bacillus amyloliquefaciens ChbB protein binds beta- and alpha-chitin and has homologues in related strains. Microbiology (Reading). 2001;147(Pt 7):1793-1803. DOI:10.1099/00221287-147-7-1793 |
- Wong E, Vaaje-Kolstad G, Ghosh A, Hurtado-Guerrero R, Konarev PV, Ibrahim AF, Svergun DI, Eijsink VG, Chatterjee NS, and van Aalten DM. (2012). The Vibrio cholerae colonization factor GbpA possesses a modular structure that governs binding to different host surfaces. PLoS Pathog. 2012;8(1):e1002373. DOI:10.1371/journal.ppat.1002373 |
- Folders J, Tommassen J, van Loon LC, and Bitter W. (2000). Identification of a chitin-binding protein secreted by Pseudomonas aeruginosa. J Bacteriol. 2000;182(5):1257-63. DOI:10.1128/JB.182.5.1257-1263.2000 |
- Vaaje-Kolstad G, Bunaes AC, Mathiesen G, and Eijsink VG. (2009). The chitinolytic system of Lactococcus lactis ssp. lactis comprises a nonprocessive chitinase and a chitin-binding protein that promotes the degradation of alpha- and beta-chitin. FEBS J. 2009;276(8):2402-15. DOI:10.1111/j.1742-4658.2009.06972.x |
- Walter S and Schrempf H. (2008). Characteristics of the surface-located carbohydrate-binding protein CbpC from Streptomyces coelicolor A32. Arch Microbiol. 2008;190(2):119-27. DOI:10.1007/s00203-008-0373-7 |
- Sánchez B, González-Tejedo C, Ruas-Madiedo P, Urdaci MC, and Margolles A. (2011). Lactobacillus plantarum extracellular chitin-binding protein and its role in the interaction between chitin, Caco-2 cells, and mucin. Appl Environ Microbiol. 2011;77(3):1123-6. DOI:10.1128/AEM.02080-10 |
- Loose JS, Forsberg Z, Fraaije MW, Eijsink VG, and Vaaje-Kolstad G. (2014). A rapid quantitative activity assay shows that the Vibrio cholerae colonization factor GbpA is an active lytic polysaccharide monooxygenase. FEBS Lett. 2014;588(18):3435-40. DOI:10.1016/j.febslet.2014.07.036 |
- Vaaje-Kolstad G, Forsberg Z, Loose JS, Bissaro B, and Eijsink VG. (2017). Structural diversity of lytic polysaccharide monooxygenases. Curr Opin Struct Biol. 2017;44:67-76. DOI:10.1016/j.sbi.2016.12.012 |
- Phillips CM, Beeson WT, Cate JH, and Marletta MA. (2011). Cellobiose dehydrogenase and a copper-dependent polysaccharide monooxygenase potentiate cellulose degradation by Neurospora crassa. ACS Chem Biol. 2011;6(12):1399-406. DOI:10.1021/cb200351y |
- Isaksen T, Westereng B, Aachmann FL, Agger JW, Kracher D, Kittl R, Ludwig R, Haltrich D, Eijsink VG, and Horn SJ. (2014). A C4-oxidizing lytic polysaccharide monooxygenase cleaving both cellulose and cello-oligosaccharides. J Biol Chem. 2014;289(5):2632-42. DOI:10.1074/jbc.M113.530196 |
- Bissaro B, Røhr ÅK, Müller G, Chylenski P, Skaugen M, Forsberg Z, Horn SJ, Vaaje-Kolstad G, and Eijsink VGH. (2017). Oxidative cleavage of polysaccharides by monocopper enzymes depends on H(2)O(2). Nat Chem Biol. 2017;13(10):1123-1128. DOI:10.1038/nchembio.2470 |
- Bertini L, Breglia R, Lambrughi M, Fantucci P, De Gioia L, Borsari M, Sola M, Bortolotti CA, and Bruschi M. (2018). Catalytic Mechanism of Fungal Lytic Polysaccharide Monooxygenases Investigated by First-Principles Calculations. Inorg Chem. 2018;57(1):86-97. DOI:10.1021/acs.inorgchem.7b02005 |
- Meier KK, Jones SM, Kaper T, Hansson H, Koetsier MJ, Karkehabadi S, Solomon EI, Sandgren M, and Kelemen B. (2018). Oxygen Activation by Cu LPMOs in Recalcitrant Carbohydrate Polysaccharide Conversion to Monomer Sugars. Chem Rev. 2018;118(5):2593-2635. DOI:10.1021/acs.chemrev.7b00421 |
- Beeson WT, Vu VV, Span EA, Phillips CM, and Marletta MA. (2015). Cellulose degradation by polysaccharide monooxygenases. Annu Rev Biochem. 2015;84:923-46. DOI:10.1146/annurev-biochem-060614-034439 |
- Walton PH and Davies GJ. (2016). On the catalytic mechanisms of lytic polysaccharide monooxygenases. Curr Opin Chem Biol. 2016;31:195-207. DOI:10.1016/j.cbpa.2016.04.001 |
- Kuusk S, Bissaro B, Kuusk P, Forsberg Z, Eijsink VGH, Sørlie M, and Väljamäe P. (2018). Kinetics of H(2)O(2)-driven degradation of chitin by a bacterial lytic polysaccharide monooxygenase. J Biol Chem. 2018;293(2):523-531. DOI:10.1074/jbc.M117.817593 |
- Agger JW, Isaksen T, Várnai A, Vidal-Melgosa S, Willats WG, Ludwig R, Horn SJ, Eijsink VG, and Westereng B. (2014). Discovery of LPMO activity on hemicelluloses shows the importance of oxidative processes in plant cell wall degradation. Proc Natl Acad Sci U S A. 2014;111(17):6287-92. DOI:10.1073/pnas.1323629111 |
- Frandsen KE, Simmons TJ, Dupree P, Poulsen JC, Hemsworth GR, Ciano L, Johnston EM, Tovborg M, Johansen KS, von Freiesleben P, Marmuse L, Fort S, Cottaz S, Driguez H, Henrissat B, Lenfant N, Tuna F, Baldansuren A, Davies GJ, Lo Leggio L, and Walton PH. (2016). The molecular basis of polysaccharide cleavage by lytic polysaccharide monooxygenases. Nat Chem Biol. 2016;12(4):298-303. DOI:10.1038/nchembio.2029 |
- Cannella D, Möllers KB, Frigaard NU, Jensen PE, Bjerrum MJ, Johansen KS, and Felby C. (2016). Light-driven oxidation of polysaccharides by photosynthetic pigments and a metalloenzyme. Nat Commun. 2016;7:11134. DOI:10.1038/ncomms11134 |
- Gudmundsson M, Kim S, Wu M, Ishida T, Momeni MH, Vaaje-Kolstad G, Lundberg D, Royant A, Ståhlberg J, Eijsink VG, Beckham GT, and Sandgren M. (2014). Structural and electronic snapshots during the transition from a Cu(II) to Cu(I) metal center of a lytic polysaccharide monooxygenase by X-ray photoreduction. J Biol Chem. 2014;289(27):18782-92. DOI:10.1074/jbc.M114.563494 |
- Quinlan RJ, Sweeney MD, Lo Leggio L, Otten H, Poulsen JC, Johansen KS, Krogh KB, Jørgensen CI, Tovborg M, Anthonsen A, Tryfona T, Walter CP, Dupree P, Xu F, Davies GJ, and Walton PH. (2011). Insights into the oxidative degradation of cellulose by a copper metalloenzyme that exploits biomass components. Proc Natl Acad Sci U S A. 2011;108(37):15079-84. DOI:10.1073/pnas.1105776108 |
- Harris PV, Welner D, McFarland KC, Re E, Navarro Poulsen JC, Brown K, Salbo R, Ding H, Vlasenko E, Merino S, Xu F, Cherry J, Larsen S, and Lo Leggio L. (2010). Stimulation of lignocellulosic biomass hydrolysis by proteins of glycoside hydrolase family 61: structure and function of a large, enigmatic family. Biochemistry. 2010;49(15):3305-16. DOI:10.1021/bi100009p |
- Hemsworth GR, Taylor EJ, Kim RQ, Gregory RC, Lewis SJ, Turkenburg JP, Parkin A, Davies GJ, and Walton PH. (2013). The copper active site of CBM33 polysaccharide oxygenases. J Am Chem Soc. 2013;135(16):6069-77. DOI:10.1021/ja402106e |
- Hemsworth GR, Davies GJ, and Walton PH. (2013). Recent insights into copper-containing lytic polysaccharide mono-oxygenases. Curr Opin Struct Biol. 2013;23(5):660-8. DOI:10.1016/j.sbi.2013.05.006 |
- Bacik JP, Mekasha S, Forsberg Z, Kovalevsky AY, Vaaje-Kolstad G, Eijsink VGH, Nix JC, Coates L, Cuneo MJ, Unkefer CJ, and Chen JC. (2017). Neutron and Atomic Resolution X-ray Structures of a Lytic Polysaccharide Monooxygenase Reveal Copper-Mediated Dioxygen Binding and Evidence for N-Terminal Deprotonation. Biochemistry. 2017;56(20):2529-2532. DOI:10.1021/acs.biochem.7b00019 |
- Courtade G, Wimmer R, Røhr ÅK, Preims M, Felice AK, Dimarogona M, Vaaje-Kolstad G, Sørlie M, Sandgren M, Ludwig R, Eijsink VG, and Aachmann FL. (2016). Interactions of a fungal lytic polysaccharide monooxygenase with β-glucan substrates and cellobiose dehydrogenase. Proc Natl Acad Sci U S A. 2016;113(21):5922-7. DOI:10.1073/pnas.1602566113 |
- Vaaje-Kolstad G, Horn SJ, van Aalten DM, Synstad B, and Eijsink VG. (2005). The non-catalytic chitin-binding protein CBP21 from Serratia marcescens is essential for chitin degradation. J Biol Chem. 2005;280(31):28492-7. DOI:10.1074/jbc.M504468200 |
- Span EA, Suess DLM, Deller MC, Britt RD, and Marletta MA. (2017). The Role of the Secondary Coordination Sphere in a Fungal Polysaccharide Monooxygenase. ACS Chem Biol. 2017;12(4):1095-1103. DOI:10.1021/acschembio.7b00016 |
- Schnellmann J, Zeltins A, Blaak H, and Schrempf H. (1994). The novel lectin-like protein CHB1 is encoded by a chitin-inducible Streptomyces olivaceoviridis gene and binds specifically to crystalline alpha-chitin of fungi and other organisms. Mol Microbiol. 1994;13(5):807-19. DOI:10.1111/j.1365-2958.1994.tb00473.x |