CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Carbohydrate Binding Module Family 81
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| CAZy DB link | |
| https://www.cazy.org/CBM81.html |
Ligand specificities
The family CBM81 was first described on September 12, 2016 [1]. According to CAZy, other members were found exclusively in gammaproteobacteria [2]. The founding CBM81 (named CBM_E1) was identified from a sugar cane soil metagenome library [3], as part of a GH5 endoglucanase, where the catalytic module and the CBM are connected by a 32 amino acids (serine-rich) linker. The CBM_E1 interaction with soluble ligands was determined by Isothermal Titration Calorimetry, resulting in the highest affinity with barley β-glucan (KA of 1.4 x 10-4 M-1), followed by cellohexaose (1.2 x 10-4 M-1), xyloglucan (0.5 x 10-4 M-1) and cellopentaose (0.4 x 10-4 M-1) [1]. The protein did not show any affinity for xylan and oligosaccharides such as cellotetraose, mannohexaose, and xylohexaose. The thermodynamic parameters indicated that the CBM_E1 binding to ligands is enthalpically driven, which is a typical characteristic of type B CBMs [4]. On the other hand, based on pull-down assays with insoluble carbohydrates, the CBM_E1 was able to bind to Avicel, but not to Bacterial Microcrystalline Cellulose (BMCC), which is characteristic of type A CBMs [1]. The Avicel is composed of about 40% of amorphous regions [5], these disordered regions of the polysaccharide are the probable CBM_E1 binding region.
Structural Features
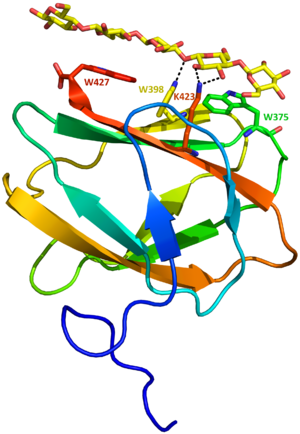
The first crystallographic structures from CBM81 family were deposited in the Protein Data Bank in 2016. Two structures from CBM_E1, in the absence (PDB ID 5KLC) and presence of cellopentaose (PDB ID 5KLE), were obtained independent of the catalytic module [1]. CBM_E1 is composed of a beta sandwich with four and five beta-strands connected by loops (Figure 1). Parallel to the beta-strands, three tryptophan are exposed to solvent forming a planar surface, which coordinates ligand binding. No differences were observed between apo and ligand complexed structures.
CBM81 is the first CBM family to exhibit mixed characteristics from type A and type B CBM binding sites. Type A CBMs bind to the surface of crystalline polysaccharides (such as cellulose and chitin) through CH-pi interactions between the surface aromatic residues and the monosaccharide units [6]. Although the planar surface of this CBM81 member is composed of aromatic residues, similar to type A CBMs, in the crystal structure of the complex one of the tryptophans has the indole ring perpendicular to the oligosaccharide chain, leading to a hydrogen bond instead of a hydrophobic interaction [1]. This observation explains the enthalpically driven binding between the CBM and the ligand. In conclusion, CBM_E1 is classified as type B, due to its affinity for soluble glycan chains instead of crystalline polysaccharides, but presents the type A flat binding-site.
Functionalities
CBM_E1 demonstrated binding to amorphous regions of cellulose, as well as beta-glucan, xyloglucan and cello-oligosaccharides [1]. All these ligands are typical substrates of endoglucanases, which are the enzymes linked to the CBM81 members so far deposited in CAZy. CBM81 may enhance endoglucanase activity by bringing the enzyme module into proximity of the substrate [7]; however, this effect has not been experimentally demonstrated for this family.
Family Firsts
- First Identified
- The CBM81 CBM_E1 was identified as part of a GH5 endoglucanase, originating from an uncultured microorganism (metagenomics) [1].
- First Structural Characterization
- The first crystal structures of the family CBM81 were from CBM_E1, in absence and presence of the ligand cellopentaose [1].
References
- Campos BM, Liberato MV, Alvarez TM, Zanphorlin LM, Ematsu GC, Barud H, Polikarpov I, Ruller R, Gilbert HJ, Zeri AC, and Squina FM. (2016). A Novel Carbohydrate-binding Module from Sugar Cane Soil Metagenome Featuring Unique Structural and Carbohydrate Affinity Properties. J Biol Chem. 2016;291(45):23734-23743. DOI:10.1074/jbc.M116.744383 |
- Weiner RM, Taylor LE 2nd, Henrissat B, Hauser L, Land M, Coutinho PM, Rancurel C, Saunders EH, Longmire AG, Zhang H, Bayer EA, Gilbert HJ, Larimer F, Zhulin IB, Ekborg NA, Lamed R, Richardson PM, Borovok I, and Hutcheson S. (2008). Complete genome sequence of the complex carbohydrate-degrading marine bacterium, Saccharophagus degradans strain 2-40 T. PLoS Genet. 2008;4(5):e1000087. DOI:10.1371/journal.pgen.1000087 |
- Alvarez TM, Paiva JH, Ruiz DM, Cairo JP, Pereira IO, Paixão DA, de Almeida RF, Tonoli CC, Ruller R, Santos CR, Squina FM, and Murakami MT. (2013). Structure and function of a novel cellulase 5 from sugarcane soil metagenome. PLoS One. 2013;8(12):e83635. DOI:10.1371/journal.pone.0083635 |
- Georgelis N, Yennawar NH, and Cosgrove DJ. (2012). Structural basis for entropy-driven cellulose binding by a type-A cellulose-binding module (CBM) and bacterial expansin. Proc Natl Acad Sci U S A. 2012;109(37):14830-5. DOI:10.1073/pnas.1213200109 |
- Hall M, Bansal P, Lee JH, Realff MJ, and Bommarius AS. (2010). Cellulose crystallinity--a key predictor of the enzymatic hydrolysis rate. FEBS J. 2010;277(6):1571-82. DOI:10.1111/j.1742-4658.2010.07585.x |
- Gilbert HJ, Knox JP, and Boraston AB. (2013). Advances in understanding the molecular basis of plant cell wall polysaccharide recognition by carbohydrate-binding modules. Curr Opin Struct Biol. 2013;23(5):669-77. DOI:10.1016/j.sbi.2013.05.005 |
- Hervé C, Rogowski A, Blake AW, Marcus SE, Gilbert HJ, and Knox JP. (2010). Carbohydrate-binding modules promote the enzymatic deconstruction of intact plant cell walls by targeting and proximity effects. Proc Natl Acad Sci U S A. 2010;107(34):15293-8. DOI:10.1073/pnas.1005732107 |