CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Carbohydrate Binding Module Family 10
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| CAZy DB link | |
| https://www.cazy.org/CBM10.html |
Ligand specificities
All the family 10 CBMs (CBM10s) are bacterial in origin. These CBMs were shown to bind to crystalline cellulose with an affinity (Ka) of ~2.5 x 105 M-1, while NMR titration experiments showed very weak binding of cellohexaoase (Ka of 1.0 x 102 M-1) [1]. No binding to other plant cell wall polysaccharides such as xylans, mannans or mixed linked β-glucans were detected [2].
Structural Features
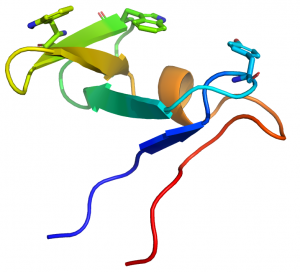
CBM10 modules generally comprise ~45 amino acids. The structure of the protein from the Cellvibrio japonicus GH10 xylanase Xyn10A was determined by NMR (see PDB ID 1E8R) [3]. The proteins displays a oligonucleotide/oligosaccharide-binding (OB) fold and is the only CBM family, to date, to adopt this secondary structure. The structure of CBM10 consists of five β-strands, organized as two antiparallel sheets, one of three strands (β-sheet 1) and one of two (β-sheet 2) that are approximately perpendicular to each other. The structure also contains a short stretch of α-helix. The protein is stabilized by two disulfide bridges.
The CBM10 module displays a planar surface that contains three aromatic residues, two contributed by the middle strand of β-sheet 1 (Trp22 and Trp24) and one by the loop connecting the strands in β-sheet 2 (Tyr8) (Fig. 1). Chemical modification of the CBM10 module with N-bromosuccinimide in combination with mutagenesis confirmed that Trp24 and Trp22 were on the surface of the protein [4]. The position of the three planar aromatic residues indicate that they make hydrophobic interactions with glucose residues n, n+2 and n+4 in the chains of crystalline cellulose. Mutagenesis studies confirmed the critical role played by the aromatic residues in cellulose recognition [4]. The importance of the three planar residues was also proposed by their conservation (tryptophans or tyrosines) in other CBM10 members. Similarly the four cysteines that formed the two disulfide bonds are invariant in the family and it was suggested that they played a generic role in protein stabilization. The planar topography of the ligand binding site and the observed specificity for insoluble cellulose indicate that the protein module is a type A CBM.
Functionalities
CBM10 modules have been observed in characterized cellulases [5], xylanases [1, 2], mannanases [6] and lytic polysaccharide monooxygenases (LPMOs) [7]. Many of these modular enzymes contain CBMs in addition to a CBM10 member, with CBM10-CBM2 modular motifs being particularly common in glycoside hydrolases from C. japonicus [8] and Saccharophagus degradans [9]. CBM10s were shown to enhance the activity of cellulases [1] and LPMOs [7] against crystalline cellulose, and it was proposed that this capacity to boost catalytic function was through increasing the local concentration or proximity of the enzyme in the vicinity of the substrate. CBM10-CBM2 combinations in single enzymes did not increase affinity through avidity effects or act synergistically to enhance cellulase activity [1]. Fluorescent imaging studies showed that the type A CBM2 and CBM10 cellulose binding modules, derived from C. japonicas Xyn10A, displayed differential binding to plant cell walls that was dependent upon cell type, tissue, and taxon of origin [10]. These modules are deployed in the exploration of the architecture of plant cell walls [10, 11].
Family Firsts
- First Identified
- CBM10 modules were first discovered in the xylanase Xyn11A (formerly XynE) from C. japonicus [2].
- First structural characterization
- The first 3D structure of a CBM10 module, which was determined by NMR, was from the C. japonicus xylanase Xyn10A [3].
References
- Gill J, Rixon JE, Bolam DN, McQueen-Mason S, Simpson PJ, Williamson MP, Hazlewood GP, and Gilbert HJ. (1999). The type II and X cellulose-binding domains of Pseudomonas xylanase A potentiate catalytic activity against complex substrates by a common mechanism. Biochem J. 1999;342 ( Pt 2)(Pt 2):473-80. | Google Books | Open Library
- Millward-Sadler SJ, Davidson K, Hazlewood GP, Black GW, Gilbert HJ, and Clarke JH. (1995). Novel cellulose-binding domains, NodB homologues and conserved modular architecture in xylanases from the aerobic soil bacteria Pseudomonas fluorescens subsp. cellulosa and Cellvibrio mixtus. Biochem J. 1995;312 ( Pt 1)(Pt 1):39-48. DOI:10.1042/bj3120039 |
- Raghothama S, Simpson PJ, Szabó L, Nagy T, Gilbert HJ, and Williamson MP. (2000). Solution structure of the CBM10 cellulose binding module from Pseudomonas xylanase A. Biochemistry. 2000;39(5):978-84. DOI:10.1021/bi992163+ |
- Ponyi T, Szabó L, Nagy T, Orosz L, Simpson PJ, Williamson MP, and Gilbert HJ. (2000). Trp22, Trp24, and Tyr8 play a pivotal role in the binding of the family 10 cellulose-binding module from Pseudomonas xylanase A to insoluble ligands. Biochemistry. 2000;39(5):985-91. DOI:10.1021/bi9921642 |
- Hall J, Black GW, Ferreira LM, Millward-Sadler SJ, Ali BR, Hazlewood GP, and Gilbert HJ. (1995). The non-catalytic cellulose-binding domain of a novel cellulase from Pseudomonas fluorescens subsp. cellulosa is important for the efficient hydrolysis of Avicel. Biochem J. 1995;309 ( Pt 3)(Pt 3):749-56. DOI:10.1042/bj3090749 |
- Hogg D, Pell G, Dupree P, Goubet F, Martín-Orúe SM, Armand S, and Gilbert HJ. (2003). The modular architecture of Cellvibrio japonicus mannanases in glycoside hydrolase families 5 and 26 points to differences in their role in mannan degradation. Biochem J. 2003;371(Pt 3):1027-43. DOI:10.1042/BJ20021860 |
- Crouch LI, Labourel A, Walton PH, Davies GJ, and Gilbert HJ. (2016). The Contribution of Non-catalytic Carbohydrate Binding Modules to the Activity of Lytic Polysaccharide Monooxygenases. J Biol Chem. 2016;291(14):7439-49. DOI:10.1074/jbc.M115.702365 |
- DeBoy RT, Mongodin EF, Fouts DE, Tailford LE, Khouri H, Emerson JB, Mohamoud Y, Watkins K, Henrissat B, Gilbert HJ, and Nelson KE. (2008). Insights into plant cell wall degradation from the genome sequence of the soil bacterium Cellvibrio japonicus. J Bacteriol. 2008;190(15):5455-63. DOI:10.1128/JB.01701-07 |
- Weiner RM, Taylor LE 2nd, Henrissat B, Hauser L, Land M, Coutinho PM, Rancurel C, Saunders EH, Longmire AG, Zhang H, Bayer EA, Gilbert HJ, Larimer F, Zhulin IB, Ekborg NA, Lamed R, Richardson PM, Borovok I, and Hutcheson S. (2008). Complete genome sequence of the complex carbohydrate-degrading marine bacterium, Saccharophagus degradans strain 2-40 T. PLoS Genet. 2008;4(5):e1000087. DOI:10.1371/journal.pgen.1000087 |
- Blake AW, McCartney L, Flint JE, Bolam DN, Boraston AB, Gilbert HJ, and Knox JP. (2006). Understanding the biological rationale for the diversity of cellulose-directed carbohydrate-binding modules in prokaryotic enzymes. J Biol Chem. 2006;281(39):29321-9. DOI:10.1074/jbc.M605903200 |
- McCartney L, Gilbert HJ, Bolam DN, Boraston AB, and Knox JP. (2004). Glycoside hydrolase carbohydrate-binding modules as molecular probes for the analysis of plant cell wall polymers. Anal Biochem. 2004;326(1):49-54. DOI:10.1016/j.ab.2003.11.011 |