CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Carbohydrate Binding Module Family 42
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| CAZy DB link | |
| https://www.cazy.org/CBM42.html |
Ligand specificities
This module was originally identified as a non-catalytic xylan-binding domain in GH54 α-L-arabinofuranosidase from Trichoderma reesei PC-3-7 [1]. In 2004, it was found to be a CBM specific for an L-arabinofuranosyl group because L-arabinofuranose molecules were bound to a non-catalytic domain of GH54 α-L-arabinofuranosidase B from Aspergillus kawachii (AkAbfB) [2]. CBM42 members have multivalent (usually divalent or trivalent) binding ability to non-reducing end L-arabinofuranosyl residues, which are present in plant polysaccharides (hemicelluloses) such as arabinoxylan, arabinan, and arabinogalactan. Most of CBM42s are associated with α-L-arabinofuranosidases from fungi (GH54) or bacteria (GH43).
Structural Features
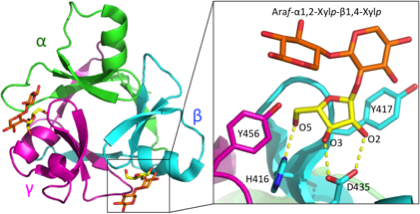
CBM42s have a β-trefoil fold that is similar to CBM13 and R(ricin)-type lectins (Figure 1). The module has a sequential 3-fold internal repeat of approximately 45 amino acid residues comprising three subdomains. The three subdomains are denoted as α, β, and γ. Each subdomain contains a discrete ligand binding site, but one of the three subdomains sometimes loses its function due to mutations at critical residues for ligand binding. CBM42s are typical Type C CBMs that bind termini of glycans with pocket-type binding sites for short oligosaccharides. The binding pockets are small but can accommodate the branched side chain L-arabinofuranosyl moiety attached to the xylan backbone of arabinoxylans [3]. The binding sites are located at side pockets of the triangular structure of the β-trefoil fold. Residues important for the binding to a non-reducing end L-arabinofuranosyl group are as follows: an aspartate forming hydrogen bonds to the O2 and O3 hydroxyls, a histidine forming a hydrogen bond to the O5 hydroxyl, and two aromatic residues (tyrosine, tryptophan, or phenylalanine) stacking to the furanose sugar (Figure 1). They are D425, H416, Y417 and Y456 in the β-subdomain of AkAbfB and are conserved in functional CBM42 subdomains.
Several crystal structures including complex structures with compounds containing an L-arabinofuranosyl group are available. For example, AkAbfB (α-subdomain is non-functional) (1wd3 1wd4) [2], (2d43 2d44) [3]; Exo-1,5-α-L-arabinofuranosidase from Sreptomyces avermitilis (all subdomains are functional) (3akf 3akg 3akh 3aki) [4]; CBM42A in Cthe_0015 from Clostridium thermocellum (β-subdomain is non-functional) (3kmv) [5].
Functionalities
CBM42s are thought to target catalytic modules (usually α-L-arabinofuranosidases) to hemicelluloses that have L-arabinofuranosyl termini or branches. Mutations at the binding sites of CBM42 significantly reduced the catalytic activity toward natural polysaccharides in these enzymes, whereas the activity toward p-nitrophenyl α-L-arabinofuranoside was not affected [3, 5]. These modules are commonly associated with α-L-arabinofuranosidases or exo-arabinanases of GH2, GH43, GH54, GH93, or non-classified GHs [5]. A survey using the GH-CBM tool shows that CBM42s are also associated with GH16 or GH30. As a typical exo-type (Type C) CBM, the CBM42 in AkAbfB shows relatively low binding affinities to L-arabinofuranose-containing oligosaccharides with Ka values of 1~5 ×103 M-1. [3]. In a novel application of CBM42 molecules, a CBM42 was fused to a feruloyl esterase to create a chimeric enzyme with enhanced thermostability and exhibited a four-fold higher activity on insoluble arabinoxylan [6].
Family Firsts
- First Identified
The L-arabinofuranose-binding function of CBM42 was first suggested by crystallography of α-L-arabinofuranosidase B from Aspergillus kawachii (AkAbfB) [2].
- First Structural Characterization
The first structure of CBM42 was revealed in 2004 in the x-ray crystal structure of AkAbfB in complex with arabinose as a full-length structure with a catalytic GH54 domain 1wd4 [2].
References
- Nogawa M, Yatsui K, Tomioka A, Okada H, and Morikawa Y. (1999). An alpha-L-arabinofuranosidase from Trichoderma reesei containing a noncatalytic xylan-binding domain. Appl Environ Microbiol. 1999;65(9):3964-8. DOI:10.1128/AEM.65.9.3964-3968.1999 |
- Miyanaga A, Koseki T, Matsuzawa H, Wakagi T, Shoun H, and Fushinobu S. (2004). Crystal structure of a family 54 alpha-L-arabinofuranosidase reveals a novel carbohydrate-binding module that can bind arabinose. J Biol Chem. 2004;279(43):44907-14. DOI:10.1074/jbc.M405390200 |
- Miyanaga A, Koseki T, Miwa Y, Mese Y, Nakamura S, Kuno A, Hirabayashi J, Matsuzawa H, Wakagi T, Shoun H, and Fushinobu S. (2006). The family 42 carbohydrate-binding module of family 54 alpha-L-arabinofuranosidase specifically binds the arabinofuranose side chain of hemicellulose. Biochem J. 2006;399(3):503-11. DOI:10.1042/BJ20060567 |
- Fujimoto Z, Ichinose H, Maehara T, Honda M, Kitaoka M, and Kaneko S. (2010). Crystal structure of an Exo-1,5-{alpha}-L-arabinofuranosidase from Streptomyces avermitilis provides insights into the mechanism of substrate discrimination between exo- and endo-type enzymes in glycoside hydrolase family 43. J Biol Chem. 2010;285(44):34134-43. DOI:10.1074/jbc.M110.164251 |
- Ribeiro T, Santos-Silva T, Alves VD, Dias FM, Luís AS, Prates JA, Ferreira LM, Romão MJ, and Fontes CM. (2010). Family 42 carbohydrate-binding modules display multiple arabinoxylan-binding interfaces presenting different ligand affinities. Biochim Biophys Acta. 2010;1804(10):2054-62. DOI:10.1016/j.bbapap.2010.07.006 |
- Koseki T, Mochizuki K, Kisara H, Miyanaga A, Fushinobu S, Murayama T, and Shiono Y. (2010). Characterization of a chimeric enzyme comprising feruloyl esterase and family 42 carbohydrate-binding module. Appl Microbiol Biotechnol. 2010;86(1):155-61. DOI:10.1007/s00253-009-2224-0 |