CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 106
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH106 | |
| Clan | none |
| Mechanism | inverting |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/GH106.html | |
Substrate specificities
The glycoside hydrolases of this family are α-L-rhamnosidases (EC 3.2.1.40), and is comprised exclusively of bacterial members. The first family GH106 enzyme characterized was Rham from Sphingomonas paucimobilis FP2001. This enzyme showed activity against p-nitrophenyl α-L-rhamnopyranoside [1]. More recently, two Bacteroides thetaiotaomicron enzymes (BT0986 and BT4145) have been characterized. These enzymes are exo-acting targeting linkages present in pectin polysaccharides. BT0986 cleaves the L-Rha-α-1,2-L-Arap linkage in the terminal region of Ccain B of rhamnogalacturonan II [2]. The enzyme BT4145 targets the L-Rha-α-1,4-D-GalA linkage in the backbone of rhamnogalacturonan I [3].
Kinetics and Mechanism
1<NMR experiments showed that GH106 members act by inverting the anomeric configuration of the glycone sugar participating in the scissle glycosidic linkage (inverting mechanism), and thus mediate bond cleave through a single displacement mechanism [3]. Additionally, GH106 enzymes are Ca2+ dependent [2]. Other ion dependent enzyme families are the exo-α-mannosidases GH38, GH47 and GH92. The structure of BT0986 in complex with D-rhamnopyranose-tetrazole indicates that catalysis is proceeds via a B2,5 transition state [2].
Catalytic Residues
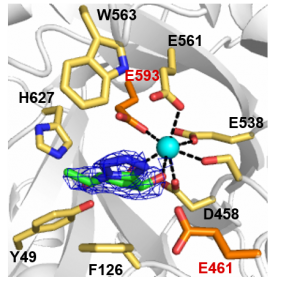
Structural characterization of B. thetaiotaomicron BT0986 identified two carboxylate amino acids (Glu461 and Glu593) that were essential for activity and are highly conserved within GH106 (Figure 1). These glutamates are separated by 8.0 Å, a distance between catalytic residues consistent with an inverting mechanism. Additionally, Glu593, which is 5.8 Å from the anomeric carbon, is ideally positioned to act as general base and Glu461 is the likely general acid as it is within hydrogen bonding distance with the glycosidic oxygen [2].
Three-dimensional structures
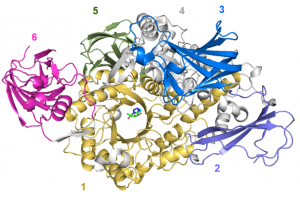
The three-dimensional structure of B. thetaiotaomicron BT0986 solved using X-ray crystallography represents the first structure of an GH106 enzyme (5MQN). BT0986 displays a N-terminal catalytic domain that presents an (β/α)8-barrel fold with several appended β-stranded domains (C-terminal) (Figure 2) [2]. Calcium makes polar interactions with O2 and O3 of rhamnose (Figure 1) and plays a role in substrate binding and the stabilization of the transition state.
Family Firsts
- First stereochemistry determination
- BT4145 from Bacteroides thetaiotaomicron [3].
- First catalytic nucleophile identification
- BT0986 from Bacteroides thetaiotaomicron [2].
- First general acid/base residue identification
- BT0986 from Bacteroides thetaiotaomicron [2].
- First 3-D structure
- BT0986 from Bacteroides thetaiotaomicron [2].
References
-
Miyata T, Kashige N, Satho T, Yamaguchi T, Aso Y and Miake F.Cloning (2005) Sequence analysis, and expression of the gene encoding Sphingomonas paucimobilis FP2001 alpha-L-rhamnosidase. Curr Microbiol, vol 51, no. 2., pp. 105-109.
- Ndeh D, Rogowski A, Cartmell A, Luis AS, Baslé A, Gray J, Venditto I, Briggs J, Zhang X, Labourel A, Terrapon N, Buffetto F, Nepogodiev S, Xiao Y, Field RA, Zhu Y, O'Neil MA, Urbanowicz BR, York WS, Davies GJ, Abbott DW, Ralet MC, Martens EC, Henrissat B, and Gilbert HJ. (2017). Complex pectin metabolism by gut bacteria reveals novel catalytic functions. Nature. 2017;544(7648):65-70. DOI:10.1038/nature21725 |
- Luis AS, Briggs J, Zhang X, Farnell B, Ndeh D, Labourel A, Baslé A, Cartmell A, Terrapon N, Stott K, Lowe EC, McLean R, Shearer K, Schückel J, Venditto I, Ralet MC, Henrissat B, Martens EC, Mosimann SC, Abbott DW, and Gilbert HJ. (2018). Dietary pectic glycans are degraded by coordinated enzyme pathways in human colonic Bacteroides. Nat Microbiol. 2018;3(2):210-219. DOI:10.1038/s41564-017-0079-1 |