CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Carbohydrate Esterase Family 15
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Carbohydrate Esterase Family CE15 | |
| Acid/alcohol sugar substrate | Acid |
| Mechanism | serine hydrolase |
| Active site residues | known, catalytic triad |
| CAZy DB link | |
| https://www.cazy.org/CE15.html | |
Substrate specificity
All CE15 enzymes characterized to date are glucuronoyl esterases (EC 3.1.1.117), cleaving esters of D-glucuronic acid. The first reported glucuronoyl esterase was ScGE1 from the white-rot fungus Schizophyllum commune, and the activity was demonstrated by TLC on a methyl ester of 4-O-methyl-D-glucuronic acid [1]. While CE15 members are found in both fungal and bacterial species, several bacterial CE15 enzymes are more promiscuous than their fungal counterparts and are active also on esters of galacturonoate [2]. Feruloyl- and acetyl esterase activities have been reported for certain CE15 enzymes as side activities [3, 4]. The proposed physiological role of CE15 enzymes is to hydrolyze lignin-carbohydrate ester linkages between lignin and glucuronoxylan in plant cell walls, and a few studies have demonstrated their activity on lignocellulose-derived materials and plant biomass [4, 5, 6].
Catalytic Residues and Mechanism
All CE15 enzymes are serine-type hydrolases, containing a catalytic triad of Glu/Asp-His-Ser [2, 7, 8, 9]. The position of the acidic residue of the triad is not similarly positioned in all CE15 members as the residue can be found on different loops of the conserved fold [9]. A conserved arginine found in all of the CE15 structures, proximal to the catalytic triad, has been proposed to stabilize the formation of the oxyanion during catalysis [2, 10, 11].
Three-dimensional structures
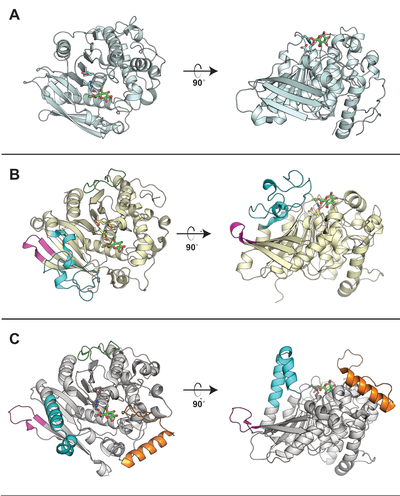
Representative structures of CE15 enzymes from bacterial and fungal sources have been determined, including TrGE (Cip2) from T. reesei (Hypocrea jecorina, PDB 3pic) [7], StGE2 from Thermothelomyces thermophila (Sporotrichum thermophile, PDB 4g4g, 4g4i, and 4g4j) [8], marine metagenome sequence MZ0003 (PDB 6ehn) [9], OtCE15A (PDB 6grw and 6gs0) and SuCE15C (PDB 6gry and 6gu8) [2] (see the CAZy database for a continuously updated list). All structurally determined CE15 enzymes share an alpha/beta hydrolase fold, consisting of a three-layer alpha-beta-alpha sandwich with the active site in a solvent-exposed cleft. The structures of the bacterial enzymes determined thus far exhibit sizeable inserts which result in much deeper active site pockets compared to the shallow active sites seen in fungal glucuronoyl esterase structures [2, 9]. The first structures with a more complex ligand than a monosaccharide were of the bacterial OtCE15A (PDB 6t0i) with a glucuronoxylooligosaccharide [10], which was followed by a similar structure of the fungal CuGE from Cerrena unicolor (PDB 6rv9).
Family Firsts
- First 3-D structure
- The first solved structure of a CE15 enzyme was the Cip2 catalytic domain from Trichoderma reesei (TrGE; PDB 3pic) [7].
- First mechanistic insight
- The crystal structure of StGE2 (from Sporotrichum thermophile) in complex with the ligand 4-O-methyl-beta-D-glucopyranuronate gave the first direct insight into substrate binding [8].
References
- Spániková S and Biely P. (2006). Glucuronoyl esterase--novel carbohydrate esterase produced by Schizophyllum commune. FEBS Lett. 2006;580(19):4597-601. DOI:10.1016/j.febslet.2006.07.033 |
- Arnling Bååth J, Mazurkewich S, Knudsen RM, Poulsen JN, Olsson L, Lo Leggio L, and Larsbrink J. (2018). Biochemical and structural features of diverse bacterial glucuronoyl esterases facilitating recalcitrant biomass conversion. Biotechnol Biofuels. 2018;11:213. DOI:10.1186/s13068-018-1213-x |
- De Santi C, Willassen NP, and Williamson A. (2016). Biochemical Characterization of a Family 15 Carbohydrate Esterase from a Bacterial Marine Arctic Metagenome. PLoS One. 2016;11(7):e0159345. DOI:10.1371/journal.pone.0159345 |
- Mosbech C, Holck J, Meyer AS, and Agger JW. (2018). The natural catalytic function of CuGE glucuronoyl esterase in hydrolysis of genuine lignin-carbohydrate complexes from birch. Biotechnol Biofuels. 2018;11:71. DOI:10.1186/s13068-018-1075-2 |
- d'Errico C, Börjesson J, Ding H, Krogh KB, Spodsberg N, Madsen R, and Monrad RN. (2016). Improved biomass degradation using fungal glucuronoyl-esterases-hydrolysis of natural corn fiber substrate. J Biotechnol. 2016;219:117-23. DOI:10.1016/j.jbiotec.2015.12.024 |
- Arnling Bååth J, Giummarella N, Klaubauf S, Lawoko M, and Olsson L. (2016). A glucuronoyl esterase from Acremonium alcalophilum cleaves native lignin-carbohydrate ester bonds. FEBS Lett. 2016;590(16):2611-8. DOI:10.1002/1873-3468.12290 |
- Pokkuluri PR, Duke NE, Wood SJ, Cotta MA, Li XL, Biely P, and Schiffer M. (2011). Structure of the catalytic domain of glucuronoyl esterase Cip2 from Hypocrea jecorina. Proteins. 2011;79(8):2588-92. DOI:10.1002/prot.23088 |
- Charavgi MD, Dimarogona M, Topakas E, Christakopoulos P, and Chrysina ED. (2013). The structure of a novel glucuronoyl esterase from Myceliophthora thermophila gives new insights into its role as a potential biocatalyst. Acta Crystallogr D Biol Crystallogr. 2013;69(Pt 1):63-73. DOI:10.1107/S0907444912042400 |
- De Santi C, Gani OA, Helland R, and Williamson A. (2017). Structural insight into a CE15 esterase from the marine bacterial metagenome. Sci Rep. 2017;7(1):17278. DOI:10.1038/s41598-017-17677-4 |
- Mazurkewich S, Poulsen JN, Lo Leggio L, and Larsbrink J. (2019). Structural and biochemical studies of the glucuronoyl esterase OtCE15A illuminate its interaction with lignocellulosic components. J Biol Chem. 2019;294(52):19978-19987. DOI:10.1074/jbc.RA119.011435 |
- Zong Z, Mazurkewich S, Pereira CS, Fu H, Cai W, Shao X, Skaf MS, Larsbrink J, and Lo Leggio L. (2022). Mechanism and biomass association of glucuronoyl esterase: an α/β hydrolase with potential in biomass conversion. Nat Commun. 2022;13(1):1449. DOI:10.1038/s41467-022-28938-w |
- Ernst HA, Mosbech C, Langkilde AE, Westh P, Meyer AS, Agger JW, and Larsen S. (2020). The structural basis of fungal glucuronoyl esterase activity on natural substrates. Nat Commun. 2020;11(1):1026. DOI:10.1038/s41467-020-14833-9 |