CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 17
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH17 | |
| Clan | GH-A |
| Mechanism | retaining |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/GH17.html | |
Substrate specificities
The family GH17 glycoside hydrolases are clan GH-A enzymes from bacteria, fungi and plants, and include two major groups of enzymes with related but distinct substrate specificities, namely 1,3-β-D-glucan endohydrolases (EC 3.2.1.39) and 1,3;1,4-β-D-glucan endohydrolases (EC 3.1.2.73). A 1,3-β-D-glucan exohydrolase (EC 3.1.2.58) is also classified in this family. The family GH17 enzymes have quite distinct amino acid sequences and 3D structures compared with the 1,3-β-D-glucan endohydrolases and 1,3;1,4-β-D-glucan endohydrolases that have similar substrate specificities but are classified in families GH16, GH55, GH64 and GH81.
The family GH17 1,3-β-D-glucan endohydrolases hydrolyse internal 1,3-β-D-glucosidic linkages in polysaccharides, but usually require a region of contiguous unbranched, un-substituted 1,3-β-D-glucosyl residues for activity. The enzymes release 1,3-β-D-oligoglucosides of DP 2-5 as their major products. Because the 1,3-β-D-glucan endohydrolases require a region of contiguous unbranched, unsubstituted 1,3-β-D-glucosyl residues for activity, they are unable to hydrolyse the single 1,3-β-D-glucosidic linkages in 1,3;1,4-β-D-glucans from the Poaceae, but they will hydrolyse 1,3-β-D-glucosidic linkages in fungal 1,3;1,6-β-D-glucans, provided an appropriate region of contiguous un-substituted 1,3-β-D-glucosyl residues is available. The family GH17 1,3;1,4-β-D-glucan endohydrolases (EC 3.1.2.73) hydrolyse 1,4-β-D-glucosidic linkages, but only 1,3;1,4-β-D-glucans in which the glucosyl residue involved in the glycosidic linkage cleaved is substituted at the C(0)3 position, that is, where the 1,4-β-D-glucosidic linkages are located on the reducing end side of 1,3-β-D-glucosyl residues.
Reaction products released are mainly 1,3;1,4-β-D-tri- and tetrasaccharides (G4G3Gred and G4G4G3Gred), but they also release higher oligosaccharides of up to 10 or more contiguous 1,4-β-D-glucosyl residues with a single reducing terminal 1,3-β-D-glucosyl residue (e.g. G4G4G4G4G4G4G3Gred). These longer oligosaccharides originate from the longer regions of adjacent 1,4-linkages that account for approximately 10% by weight of 1,3;1,4-β-D-glucans in cell walls of the Poaceae [1].
Kinetics and Mechanism
The stereochemistry of the reaction has been determined experimentally and catalysis by GH17 enzymes occurs via a classical Koshland retaining mechanism with the β-anomeric configuration of the released oligosaccharide being retained [2]. Detailed kinetic analyses are available for three purified barley 1,3-β-D-glucan endohydrolases and two barley 1,3;1,4-β-D-glucan endohydrolases [2].
Catalytic Residues
Active site labelling with epoxyalkyl β-D-oligoglucoside inhibitors identified Glu231 and Glu232 as the catalytic nucleophiles of the barley 1,3- and 1,3;1,4-β-D-glucan endohydrolases, respectively [3], located at the bottom of, and about two-thirds of the way along the substrate binding cleft. The general acid/base residue of 1,3;1,4-β-D-glucan endohydrolase was initially identified as Glu288 by chemical labelling procedures [3, 4], but this assignment was subsequently revised and Glu93 was proposed based on primary and tertiary structure similarity of GH17 enzymes with clan GH-A β-glycosidases [5, 6]. The 5-6 Å distance between Glu232 and Glu93 is typical of retaining enzymes.
Three-dimensional structures
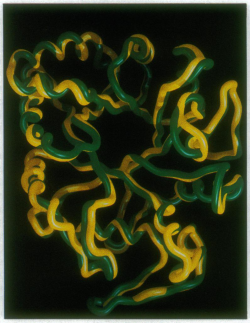
The crystal structures of the barley 1,3-β-D-glucan endohydrolase isoenzyme GII and 1,3;1,4-β-D-glucan endohydrolase isoenzyme EII have been solved to 2.2-2.3 Å resolution and shown to adopt essentially identical (β/α)8 TIM barrel structures [7]. The rms deviation in Cα positions between the two barley enzymes is 0.65 Å for 278 residues [7]. This indicates that only minor differences in structure and amino acid dispositions at the substrate-binding and catalytic sites are sufficient to change a pre-existing 1,3-β-D-glucan endohydrolase into a highly specific 1,3;1,4-β-D-glucan endohydrolase [7].
A deep substrate-binding cleft extends across the surface of the enzyme and can accommodate 6-8 glucosyl-binding subsites [7]. The open cleft enables the enzyme to bind at essentially any position along the 1,3;1,4-β-D-glucan substrate and hence to hydrolyse internal glycoside linkages. Like in other clan GH-A structures, the general acid/base and catalytic nucleophile glutamates are positioned on strands β-4 and β-7 [5, 6, 7].
The X-ray crystallographic data provide compelling evidence that the 1,3;1,4-β-D-glucan endohydrolases of barley evolved via the recruitment of pre-existing and widely distributed family GH17 1,3-β-D-glucan endohydrolases [6]. The similarities in substrate specificity between family GH17 and GH16 enzymes has arisen through convergent evolution; the family GH16 enzymes are members of clan-B and have a β-jelly roll structure.
Family Firsts
- First sterochemistry determination
- The stereochemical course of hydrolysis of Laminaria digitata laminarin and barley 1,3;1,4-β-glucan by barley 1,3-β-glucanase isoenzyme GII and 1,3;1,4-β-glucanase isoenzyme EII, respectively, was determined by 1H-NMR. Both enzymes catalyze hydrolysis with retention of anomeric configuration (e-->e) and may therefore operate via a double displacement mechanism [2].
- First catalytic nucleophile identification
- Active site labelling with epoxyalkyl-β-D-oligoglucoside inhibitors identified Glu231 and Glu232 as the catalytic nucleophiles of the barley 1,3- and 1,3;1,4-β-D-glucan endohydrolases, respectively [3].
- First general acid/base residue identification
- This residue was identified as the conserved Glu residue at the C-terminal end of strand β-4 based on primary and tertiary structure similarity of GH17 enzymes with clan GH-A β-glycosidases [5, 6].
- First 3-D structure
- Barley 1,3-β-D-glucan endohydrolase isoenzyme GII and 1,3;1,4-β-D-glucan endohydrolase isoenzyme EII, solved to 2.2-2.3 Å resolution [7].
References
-
Woodward, J.R. and Fincher, G.B. (1982) Substrate specificities and kinetic properties of two (1-3),(1-4)-β-D-glucan endo-hydrolases from germinating barley (Hordeum vulgare). Carbohydr. Res. 106, 111-122. DOI:10.1016/S0008-6215(00)80737-5
- Chen L, Sadek M, Stone BA, Brownlee RT, Fincher GB, and Høj PB. (1995). Stereochemical course of glucan hydrolysis by barley (1-->3)- and (1-->3, 1-->4)-beta-glucanases. Biochim Biophys Acta. 1995;1253(1):112-6. DOI:10.1016/0167-4838(95)00157-p |
- Chen L, Fincher GB, and Høj PB. (1993). Evolution of polysaccharide hydrolase substrate specificity. Catalytic amino acids are conserved in barley 1,3-1,4- and 1,3-beta-glucanases. J Biol Chem. 1993;268(18):13318-26. | Google Books | Open Library
- Chen L, Garrett TP, Fincher GB, and Høj PB. (1995). A tetrad of ionizable amino acids is important for catalysis in barley beta-glucanases. J Biol Chem. 1995;270(14):8093-101. DOI:10.1074/jbc.270.14.8093 |
- Pickersgill R, Harris G, Lo Leggio L, Mayans O, and Jenkins J. (1998). Superfamilies: the 4/7 superfamily of beta alpha-barrel glycosidases and the right-handed parallel beta-helix superfamily. Biochem Soc Trans. 1998;26(2):190-8. DOI:10.1042/bst0260190 |
- Henrissat B, Callebaut I, Fabrega S, Lehn P, Mornon JP, and Davies G. (1995). Conserved catalytic machinery and the prediction of a common fold for several families of glycosyl hydrolases. Proc Natl Acad Sci U S A. 1995;92(15):7090-4. DOI:10.1073/pnas.92.15.7090 |
- Varghese JN, Garrett TP, Colman PM, Chen L, Høj PB, and Fincher GB. (1994). Three-dimensional structures of two plant beta-glucan endohydrolases with distinct substrate specificities. Proc Natl Acad Sci U S A. 1994;91(7):2785-9. DOI:10.1073/pnas.91.7.2785 |
- Hrmova M and Fincher GB. (2001). Structure-function relationships of beta-D-glucan endo- and exohydrolases from higher plants. Plant Mol Biol. 2001;47(1-2):73-91. | Google Books | Open Library