CAZypedia needs your help! We have many unassigned GH, PL, CE, AA, GT, and CBM pages in need of Authors and Responsible Curators.
Scientists at all career stages, including students, are welcome to contribute to CAZypedia. Read more here, and in the 10th anniversary article in Glycobiology.
New to the CAZy classification? Read this first.
*
Consider attending the 15th Carbohydrate Bioengineering Meeting in Ghent, 5-8 May 2024.
Carbohydrate Binding Module Family 77
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| CAZy DB link | |
| http://www.cazy.org/CBM77.html |
Ligand specificities
CBM77 is a bacterial family of binding modules that comprises around 110 amino acids. Only the founding member of this family, CBM77RfPL1/9 has been characterized. This carbohydrate-binding module found in Ruminococcus flavefaciens binds exclusively to homogalacturonan (pectin) with low degrees of methyl esterification (DE) [1]. By isothermal titration calorimetry, CBM77RfPL1/9 displays high affinity for low DE pectins such as, lime DE 11% and citrus pectin DE 30% (KA 1.8 x 105 M-1 and 1.1 x 104 M-1, respectively). This binding module displays a reduction in affinity as the DE of the ligand increase. Indeed CBM77RfPL1/9 exhibits no binding to polygalacturonic acid with a DE ≥80%. This binding module displays optimum binding to oligosaccharides of α-1,4-D-Galacturonic acid (GalA) with a degree of polymerization of 7 to 8. Additionally, the presence of a chelating agent (EDTA) did not influence CBM77RfPL1/9 affinity indicating a divalent cation independent binding mechanism [1]. The only other example of a CBM that binds to homogalacturonan backbone is CBM32 from Yersinia enterolitica (YeCBM32) (which is not a component of an enzyme) [2].
Structural Features
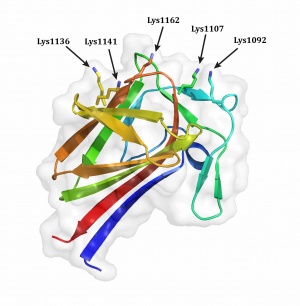
The three-dimensional structure of Ruminoccocus CBM77RfPL1/9 (5FU5) solved using X-ray displays a beta-sandwich fold in which the 13 antiparallel β-strands are organized in two β-sheets [1].
The key residues implicated in ligand binding were identified by site-direct mutagenesis. The canonical ligand binding site of an endo-binding type B CBM adopts a cleft-like structure. However, in CBM77RfPL1/9, the alanine mutation of aromatic or basic amino acids in the concave surface had no effect on ligand binding. Only the alanine mutation of three invariant basic residues (Lys1092, Lys1107, and Lys1162) located at the convex surface of CBM77RfPL1/9 abrogated binding to polygalacturonic acid (PGA), showing the location of the binding site. The single alanine mutants of two distal basic residues (Lys1136 and Lys1141) displayed only a modest reduction in affinity for PGA when compared with the wild-type protein; however, the double mutation Lys1136A/Lys1141A resulted in complete loss of binding indicating that these lysines display functional redundancy. Additionally, based on the mutant binding data, the CBM77RfPL1/9 ligand recognition surface is ~ 25 Å suggesting that the binding site can accommodate a hexasaccharide [1].
The CBM77RfPL1/9 ligand recognition mediated by basic polar residues is distinct from other CBMs where aromatic residues play a central role in glycan recognition, but resembles glycosaminoglycan binding proteins, where ligand recognition is also mediated by basic residues (arginine and lysines) [3].
Functionalities
Ruminoccocus flavefaciens CBM77RfPL1/9 is a component of an enzyme that contains two catalytic modules, belonging to polysaccharide lyase families 1 (PL1) and 9 (PL9) and a C-terminal dockerin module. Consistent with the CBM77RfPL1/9 specificity, both PL enzymes display pectate lyase activity [1].
This CBM family contains 26 members from a range of bacteria suggesting a general role in pectin degradation that is not specific to the cellulosome organization or the rumen environment. In the majority of the proteins, the CBM77 is appended to PL1 catalytic modules although PL9 sequences are also present [1].
Additionally, the binding of CBM77RfPL1/9 was also tested in the context of plant cell wall. This binding module specifically labelled homogalacturonan in Tobacco stem sections [1].
Family Firsts
- First Identified
- CBM77RfPL1/9 from Ruminococcus flavefaciens [1].
- First Structural Characterization
- The first 3D crystal structure solved was CBM77RfPL1/9 [1].
References
- Venditto I, Luis AS, Rydahl M, Schückel J, Fernandes VO, Vidal-Melgosa S, Bule P, Goyal A, Pires VM, Dourado CG, Ferreira LM, Coutinho PM, Henrissat B, Knox JP, Baslé A, Najmudin S, Gilbert HJ, Willats WG, and Fontes CM. (2016). Complexity of the Ruminococcus flavefaciens cellulosome reflects an expansion in glycan recognition. Proc Natl Acad Sci U S A. 2016;113(26):7136-41. DOI:10.1073/pnas.1601558113 |
- Abbott DW, Hrynuik S, and Boraston AB. (2007). Identification and characterization of a novel periplasmic polygalacturonic acid binding protein from Yersinia enterolitica. J Mol Biol. 2007;367(4):1023-33. DOI:10.1016/j.jmb.2007.01.030 |
- Hileman RE, Fromm JR, Weiler JM, and Linhardt RJ. (1998). Glycosaminoglycan-protein interactions: definition of consensus sites in glycosaminoglycan binding proteins. Bioessays. 1998;20(2):156-67. DOI:10.1002/(SICI)1521-1878(199802)20:2<156::AID-BIES8>3.0.CO;2-R |