CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 89
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
| Glycoside Hydrolase Family GH89 | |
| Clan | none |
| Mechanism | Retaining |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/GH89.html | |
Substrate specificities
Family 89 glycoside hydrolases are α-N-acetylglucosaminidases [1, 2]. CpGH89, produced by the animal pathogen Clostridium perfringens, is active on the unusual α-D-GlcNAc-(1,4)-D-Galactose motif [3] displayed on the gastric mucosal class III mucins [4]. The human lysosomal enzyme, NAGLU, is involved in the degradation of heparan sulfate [5, 6]. Mutations in this enzyme can cause a devastating lysosomal storage disorder called Sanfilippo syndrome type B which is also called mucopolysaccharidosis IIIB (MPS IIIB) [5, 6].
Kinetics and Mechanism
Mechanistic data is available on CpGH89, a family 89 glycoside hydrolase produced by Clostridium perfringens. CpGH89 uses a classical Koshland retaining mechanism to hydrolyze the glycosidic bond [1]. This results in hydrolysis with a net retention of stereochemistry at the anomeric carbon [1].
Catalytic Residues
Two catalytically important glutamate residues have been identified in CpGH89, Glu483 and Glu601 [1]. These residues are between 6.1-6.7Å apart, which is consistent with a retaining catalytic mechanism. Mutation of Glu601 to an alanine results in an apparent abolishment of activity suggesting this residue is active as the catalytic nucleophile. Glu601 resides below the A-face of the sugar ring and is 2.8-3.1Å from C1 and appears suitably placed for nucleophilic attack on the anomeric carbon. Mutation of Glu483 to alanine results in much less severe impairments in catalysis suggesting this residue is active as the general acid/base residue. Glu483 is ~3.6Å from C1 and appears to be positioned in such a way that it would be capable of forming a hydrogen bond with the glycosidic oxygen of the substrate.
Three-dimensional structures
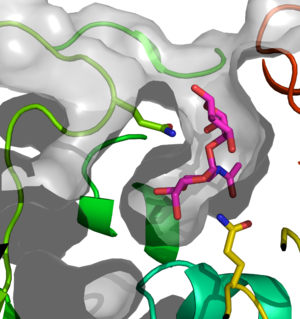
The three dimensional structure was first described for CpGH89, see PDB entries 2vc9, 2vca, 2vcb,2vcc [1] followed by the structure of a CpGH89 double catalytic residue mutant in complex with the class III mucosal disaccharide α-D-GlcNAc-(1,4)-D-Galactose, 4a4a (Figure 1) [7]. The structure of the human enzyme NAGLU, 4xwh, was released in 2019 [8].
CpGH89 is a multi-modular protein and quite large (2095 amino acids) and contains six modules with sequence similarity to the family 32 carbohydrate-binding modules (CBM32s) [9]. To obtain the CpGH89 catalytic module structure, only residues 26-916 were crystallized [1]. The N-terminal domain (residues 26-155) forms a β-sandwich fold and is one of the CBM32-like domains. This module is tightly packed against the rest of the protein through a number of hydrophobic and hydrogen bonding interactions and is not predicted to be functional in carbohydrate-binding [9]. The catalytic region is comprised of a small mixed α/β domain (residues 170-280), a decorated (α/β)8 core (residues 280-620), and an all α-helical domain (residues 621-916) [1]. The active site is sock-shaped to accommodate the bend in the class III mucosal α-D-GlcNAc-(1,4)-D-Galactose motif caused by the alpha glycosidic linkage (Figure 1) [1, 7]. The mutations causing MPS IIIB were mapped onto the bacterial homologue, demonstrating that the mutations are mainly structural in nature [1].
NAGLU has only 28.2% sequence identity with CpGH89 and lacks the CBM32-like domain. Otherwise the tertiary structures of CpGH89 and NAGLU are similar [8]. The predicted catalytic glutamates, E316 and E446, are present in NAGLU [8]. NAGLU has six potential N-glycosylation sites and appears to be N-glycosylated [8]. The recombinant enzyme was produced as a trimer in solution though the physiological relevance of the trimer is unknown [8]. In comparison, CpGH89 crystallized as a monomer in the asymmetric unit [1]. The substrate binding pocket of NAGLU is more open when compared to the sock-shaped active site of CpGH89 [1, 8]. The mutations causing MPS IIIB were also mapped onto NAGLU, confirming that the mutations are mainly structural in nature [8].
Family Firsts
- First sterochemistry determination
- 1H NMR spectroscopy reveals that CpGH89 acts with retention of stereochemistry [1].
- First catalytic nucleophile identification
- The catalytic nucleophile was revealed by site directed mutagenesis on CpGH89 Glu601 [1].
- First general acid/base residue identification
- The general acid/base was revealed by site directed mutagenesis on CpGH89 Glu483 [1].
- First 3-D structure
- See PDB entries 2vc9, 2vca, 2vcb and 2vcc [1].
References
- Ficko-Blean E, Stubbs KA, Nemirovsky O, Vocadlo DJ, and Boraston AB. (2008). Structural and mechanistic insight into the basis of mucopolysaccharidosis IIIB. Proc Natl Acad Sci U S A. 2008;105(18):6560-5. DOI:10.1073/pnas.0711491105 |
- Weber B, Hopwood JJ, and Yogalingam G. (2001). Expression and characterization of human recombinant and alpha-N-acetylglucosaminidase. Protein Expr Purif. 2001;21(2):251-9. DOI:10.1006/prep.2000.1361 |
- Fujita M, Tsuchida A, Hirata A, Kobayashi N, Goto K, Osumi K, Hirose Y, Nakayama J, Yamanoi T, Ashida H, and Mizuno M. (2011). Glycoside hydrolase family 89 alpha-N-acetylglucosaminidase from Clostridium perfringens specifically acts on GlcNAc alpha1,4Gal beta1R at the non-reducing terminus of O-glycans in gastric mucin. J Biol Chem. 2011;286(8):6479-89. DOI:10.1074/jbc.M110.206722 |
- Ishihara K, Kurihara M, Goso Y, Urata T, Ota H, Katsuyama T, and Hotta K. (1996). Peripheral alpha-linked N-acetylglucosamine on the carbohydrate moiety of mucin derived from mammalian gastric gland mucous cells: epitope recognized by a newly characterized monoclonal antibody. Biochem J. 1996;318 ( Pt 2)(Pt 2):409-16. DOI:10.1042/bj3180409 |
- Yogalingam G and Hopwood JJ. (2001). Molecular genetics of mucopolysaccharidosis type IIIA and IIIB: Diagnostic, clinical, and biological implications. Hum Mutat. 2001;18(4):264-81. DOI:10.1002/humu.1189 |
- Weber B, Guo XH, Kleijer WJ, van de Kamp JJ, Poorthuis BJ, and Hopwood JJ. (1999). Sanfilippo type B syndrome (mucopolysaccharidosis III B): allelic heterogeneity corresponds to the wide spectrum of clinical phenotypes. Eur J Hum Genet. 1999;7(1):34-44. DOI:10.1038/sj.ejhg.5200242 |
- Ficko-Blean E and Boraston AB. (2012). Structural analysis of a bacterial exo-α-D-N-acetylglucosaminidase in complex with an unusual disaccharide found in class III mucin. Glycobiology. 2012;22(5):590-5. DOI:10.1093/glycob/cwr165 |
- Birrane G, Dassier AL, Romashko A, Lundberg D, Holmes K, Cottle T, Norton AW, Zhang B, Concino MF, and Meiyappan M. (2019). Structural characterization of the α-N-acetylglucosaminidase, a key enzyme in the pathogenesis of Sanfilippo syndrome B. J Struct Biol. 2019;205(3):65-71. DOI:10.1016/j.jsb.2019.02.005 |
- Ficko-Blean E, Stuart CP, Suits MD, Cid M, Tessier M, Woods RJ, and Boraston AB. (2012). Carbohydrate recognition by an architecturally complex α-N-acetylglucosaminidase from Clostridium perfringens. PLoS One. 2012;7(3):e33524. DOI:10.1371/journal.pone.0033524 |