CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article.
Totally new to the CAZy classification? Read this first.
Glycosyltransferase Family 42
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Warren Wakarchuk^^^
- Responsible Curator: ^^^Warren Wakarchuk^^^
| Glycosyltransferase Family GT42 | |
| Clan | GT-x |
| Mechanism | inverting, donor is CMP-beta-Neu5Ac |
| Active site residues | His188 is the catalytic base in the Campylobacter jejuni GT-42 known as Cst-II
His201 is the catlytic base in theCampylobacter jejuniGT42 known as Cst-I |
| CAZy DB link | |
| http://www.cazy.org/fam/GT42.html | |
Acceptor specificities
The enzymes in GT-42 were originally examined from isolates of C. jejuni that express a number of ganglioside mimics, some of which have an a2,8-a2,3 linked di-sialic acid moiety [1]. The CjGT-42 known as Cst-II was found as part of the LOS biosynthesis operon [2], which facilitated the correlation of gene content to LOS structure. Based on the LOS structures which contained both a-2,3 and a-2,8 linked sialic acid it was predicted that there should be two sialyltransferases, but the surprising finding was that a single CjGT-42 enzyme, Cst-II, was making both linkages [3]. This bi-functional enzyme has been shown to be a determinant in the development of post-infectious neuropathies (Guillain-Barré syndrome), due to an anti-ganglioside antibody response in some patients who were infected with C. jejuni [4] [5]. Some species of Campylobacter express a second GT-42 enzyme known as Cst-I, which was in fact cloned first, but it has been shown not to be involved in LOS biosynthesis on the basis of gene knockouts in strains with both cst genes (Michel Gilbert, personal communication). Cst-I contains an extended C-terminal domain of unknown function, and the target acceptor of this second CjGT-42 is not known at present. The GT-42 family also contains members from H. influenzae and P. multocida, in which the GT-42 enzymes are one of two or three resident sialyltransferases. In H. influenzae the first GT-42 enzyme to be described was the a-2,3-sialyltransferase Lic3A [6], followed by a second a-2,3/2,8-bi-functional version known as Lic3B [7]. In H. influenzae the in vivo acceptor for Lic3A/B is the simple disaccharide lactose in contrast to the more complex ganglioside mimics seen in C. jejuni [8].
Kinetics and Mechanism
Detailed kinetic studies have revealed that this enzyme follows an unusual steady state iso ordered bi bi kinetic mechanism for carrying out sialyl transfer with inversion [9]. His188 appears to be the catalytic base, which is somewhat unusual since this role is generally performed by Glu or Asp residues. Direct measurement of its pKa by NMR showed the pKa agrees with that deduced from the pH dependence of this reaction. This required the generation of an active monomeric form of the tetrameric enzyme [9]. Chemical rescue of the H188A mutant was performed with nucleophilic anions adding weight to the assignment of the function, and further supporting the SN2-like inverting mechanism.
Catalytic Residues
His188 in CjGT-42 Cst-II from strain OH4383 appears to be the catalytic base.
Three-dimensional structures
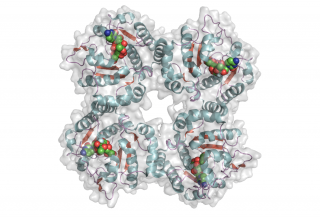
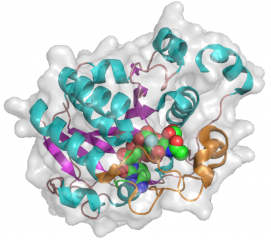
The CjGT-42 enzymes Cst-I and Cst-II were the first sialyltransferases whose structure was determined by X-ray crystallography [10][11], and they share the same basic structure. GT-42 enzymes have a modified GT-A type fold characterized by a single a/b Rossmann fold nucleotide binding domain, and a flexible lid domain composed of a coil and two helices. Unlike the typical GT-A fold, CjGT-42 enzymes lack the conserved DXD motif and are metal-ion independent. The enzyme structure revealed a tetramer in which each monomer carries an independent active site, and examination of the active site residues suggested a conserved histidine, His188 in Cst-II, as the catalytic base.
Glycosyltransferase structures often have a “lid” domain, which is a mobile loop that undergoes a conformational change upon binding a substrate [12]. The lid domain in Cst-I and Cst-II contains residues which interact with the donor and/or acceptor. A common feature of transferase substrate binding is the role of aromatic residues stacking onto the base of the sugar-nucleotide. In Cst-I the aromatic residue is Tyr171 and in Cst-II it is Tyr156, both of which are conserved in GT-42 and occur in the lid domain. There is no structure of a ternary complex for GT-42 so it is not possible to know all of the interactions that are involved in substrate binding, but there are donor complexes with the incompetent donor CMP-3-fluoro-Neu5Ac and acceptor models have been examined to predict interactions [11]. In Cst-I, Phe190 appears to prevent sialylated acceptors from binding in the active site. In Cst-II, this Phe residue is replaced by Ala175, which opens up a pocket for the carboxyl group of a modelled sialyl-lactose acceptor [11].
Family Firsts
- First general acid/base residue identification
Cite some reference here, with a short (1-2 sentence) explanation
- First 3-D structure
The CjGT-42 enzymes Cst-I and Cst-II were the first sialyltransferases whose structure was determined by X-ray crystallography [10] [11], and they share the same basic structure. GT-42 enzymes have a modified GT-A type fold characterized by a single a/b Rossmann fold nucleotide binding domain, and a flexible lid domain composed of a coil and two helices.
- PDB ID 1RO8 from GT42 (click images for large versions)
References
- Irving Nachamkin, Christine M. Szymanski, and Martin J. Blaser. (2008) Campylobacter. Amer Society for Microbiology.
- Gilbert M, Brisson JR, Karwaski MF, Michniewicz J, Cunningham AM, Wu Y, Young NM, and Wakarchuk WW. (2000). Biosynthesis of ganglioside mimics in Campylobacter jejuni OH4384. Identification of the glycosyltransferase genes, enzymatic synthesis of model compounds, and characterization of nanomole amounts by 600-mhz (1)h and (13)c NMR analysis. J Biol Chem. 2000;275(6):3896-906. DOI:10.1074/jbc.275.6.3896 |
- Gilbert M, Karwaski MF, Bernatchez S, Young NM, Taboada E, Michniewicz J, Cunningham AM, and Wakarchuk WW. (2002). The genetic bases for the variation in the lipo-oligosaccharide of the mucosal pathogen, Campylobacter jejuni. Biosynthesis of sialylated ganglioside mimics in the core oligosaccharide. J Biol Chem. 2002;277(1):327-37. DOI:10.1074/jbc.M108452200 |
- Koga M, Takahashi M, Masuda M, Hirata K, and Yuki N. (2005). Campylobacter gene polymorphism as a determinant of clinical features of Guillain-Barré syndrome. Neurology. 2005;65(9):1376-81. DOI:10.1212/01.wnl.0000176914.70893.14 |
- Hood DW, Cox AD, Gilbert M, Makepeace K, Walsh S, Deadman ME, Cody A, Martin A, Månsson M, Schweda EK, Brisson JR, Richards JC, Moxon ER, and Wakarchuk WW. (2001). Identification of a lipopolysaccharide alpha-2,3-sialyltransferase from Haemophilus influenzae. Mol Microbiol. 2001;39(2):341-50. DOI:10.1046/j.1365-2958.2001.02204.x |
- Chiu CP, Watts AG, Lairson LL, Gilbert M, Lim D, Wakarchuk WW, Withers SG, and Strynadka NC. (2004). Structural analysis of the sialyltransferase CstII from Campylobacter jejuni in complex with a substrate analog. Nat Struct Mol Biol. 2004;11(2):163-70. DOI:10.1038/nsmb720 |
- Chiu CP, Lairson LL, Gilbert M, Wakarchuk WW, Withers SG, and Strynadka NC. (2007). Structural analysis of the alpha-2,3-sialyltransferase Cst-I from Campylobacter jejuni in apo and substrate-analogue bound forms. Biochemistry. 2007;46(24):7196-204. DOI:10.1021/bi602543d |
- Lairson LL, Henrissat B, Davies GJ, and Withers SG. (2008). Glycosyltransferases: structures, functions, and mechanisms. Annu Rev Biochem. 2008;77:521-55. DOI:10.1146/annurev.biochem.76.061005.092322 |
- van Belkum A, van den Braak N, Godschalk P, Ang W, Jacobs B, Gilbert M, Wakarchuk W, Verbrugh H, and Endtz H. (2001). A Campylobacter jejuni gene associated with immune-mediated neuropathy. Nat Med. 2001;7(7):752-3. DOI:10.1038/89831 |
- Fox KL, Cox AD, Gilbert M, Wakarchuk WW, Li J, Makepeace K, Richards JC, Moxon ER, and Hood DW. (2006). Identification of a bifunctional lipopolysaccharide sialyltransferase in Haemophilus influenzae: incorporation of disialic acid. J Biol Chem. 2006;281(52):40024-32. DOI:10.1074/jbc.M602314200 |
- Schweda EK, Richards JC, Hood DW, and Moxon ER. (2007). Expression and structural diversity of the lipopolysaccharide of Haemophilus influenzae: implication in virulence. Int J Med Microbiol. 2007;297(5):297-306. DOI:10.1016/j.ijmm.2007.03.007 |
- Chan PH, Lairson LL, Lee HJ, Wakarchuk WW, Strynadka NC, Withers SG, and McIntosh LP. (2009). NMR spectroscopic characterization of the sialyltransferase CstII from Campylobacter jejuni: histidine 188 is the general base. Biochemistry. 2009;48(47):11220-30. DOI:10.1021/bi901606n |