CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article.
Totally new to the CAZy classification? Read this first.
Carbohydrate-binding modules
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Alicia Lammerts van Bueren^^^
- Responsible Curator: ^^^Al Boraston^^^ and ^^^Spencer Williams^^^
This page is under construction. In the meantime, please see these references for an essential introduction to the CAZy classification system: [1, 2]. CBMs, in particular, have been extensively reviewed[3, 4, 5, 6].
Overview
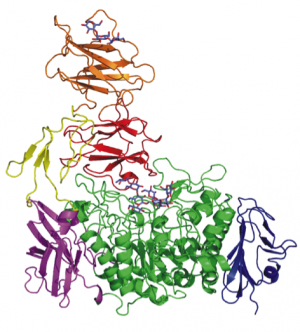
Carbohydrate-binding modules (CBMs) are defined as a stretch of amino acid sequence within a larger encoded protein sequence (Figure 1) and fold into a discreet and independent module, forming part of a larger multi-modular protein (Figure 2).
Their role is to bind to carbohydrate ligand and direct the catalytic machinery onto its substrate, thus enhancing the catalytic efficiency of the multimodular carbohydrate-active enzyme. CBMs are themselves devoid of any catalytic activity. CBMs are most commonly associated with Glycoside Hydrolases but have also been identified in Polysaccharide Lyases, polysaccharide oxidases, Glycosyltransferases and expansins.
CBMs do not undergo any conformational changes. The topography of their binding site is "preformed" to be complementary to the shape of their target carbohydrate ligand. Amino acid side chains and loops within the CBM binding pocket create the complementary ligand binding "platform" (see Types). However whole enzymes may experience complete conformational changes when binding substrate. Flexible loop regions between adjacent modules can allow for shifts in the orientation and direction of the catalytic module with respect to the CBM on the target substrate [10].
History of CBMs
CBMs were initially characterized as cellulose binding domains (CBDs) in cellobiohydrolases CBHI and CBHII from Trichoderma reesei [11, 12] and cellulases CenA and CexA from Cellulomonas fimi [13]. Limited proteolysis experiments of these enzymes yielded truncated enzyme products that showed a reduced or complete loss in their ability to hydrolyze cellulose substrates. The reduction in enzymatic activity was attributed to the loss of ~100 amino acid C-terminal domains which prevented the adsorbption of the enzymes onto cellulose substrate. Thus it was proposed that these independent "domains" are critical for targeting the enzymes onto its substrate and enhancing their hydrolytic activity.
CBDs were first organized into 13 Types based on amino acid sequence similarities [14]. This classification became complicated when similar functional domains were discovered in non-cellulolytic carbohydrate-active enzymes and did not bind cellulose (for example chitin, xylan, alpha-glucans) but met all of the criteria of CBDs. The term carbohydrate-binding module was proposed to solve this problem and encompasses all ancillary modules with non-catalytic carbohydrate-binding function (for a review see [3].
Classification
Sequence Based Classification
Carbohydrate-binding modules are currently classified into 67 families based on amino acid sequence similarities (May 2013), which are available through the Carbohydrate Active enZyme database. Sequence-based relationships often cluster together modules with similar structural folds and carbohydrate-binding function. While this is true for most CBM families, there are several families that exhibit diversity in the carbohydrate ligands they target (examples include CBM6, CBM32)
Fold
Structural information for over 90% of the CBM families is known. The most common fold exhibited by CBMs is the beta-sandwich fold which is comprised of two overlapping beta-sheets consisting of 3 - 6 antiparallel beta strands. The binding sites for carbohydrates have been shown to be located either on the same face of a beta-sheet (ref), on the edge of the beta-sheet within the joining loops (ref), or both (ref)
Picture
Other folds include the beta-trefoil fold, cysteine knot, OB fold, the hevein and hevein-like and uniqu folds [3]
Types
CBMs are classified into three main Types defined by the shape and degree of polymerization of their ligand target:
- Type A: polycrystalline surface binding
- Type B: oligosaccharides with DP>4 (mainly endo, within polysaccharide chains)
- Type C: lectin-like mono/di/tri saccharides (mainly exo, reducing/non-reducing end)
Properties of Carbohydrate Binding Interactions
Functional Roles of CBMs
CBMs carry out four main functional roles:
Targeting:
Proximity:
Cell Wall anchoring: CBM35 modules have been shown to interact with the surface glucuronic acid containing sugars in Amycolatopsis orientalis
Disruptive role has previously been described for cellulose binding CBM2a. Additionally starch binding CBM20 may have a disruptive role in amylose while dual-associated CBM41 modules may have a disruptive role in degrading glycogen granules. CBM33 was thought to have a disruptive effect on chitin, however these have now been reclassified as lytic oxygenases (expand).
Driving Forces of CBM/Carbohydrate Interactions
There are two key features that drive CBM/carbohydrate interactions. Extensive hydrogen bonding occurs between the hydroxyl groups of carbohydrate ligands and polar amino acid residues within the binding site. Additional water-mediated hydrogen bonding networks between these groups can also be found in the binding site. By far the most important characteristic driving force mediating protein-carbohydrate interactions is the position and orientation of aromatic amino acid residues (Try, Tyr and sometimes Phe) within the binding site. These essential planar residues form hydrophobic stacking interactions with the planar face of sugar rings. Weak intermolecular electrostatic interactions occur between C-H and pi electrons in the planar ring systems and contribute 1.5 - 2.5 kcal/mol energy to the binding reaction [15].
CBMs and Multivalency
CBMs and Lectins
Criteria for Defining a new CBM family
In order to define a new CBM family, one must:
- Demonstrate an independent module as part of a larger carbohydrate-active enzyme.
- Demonstrate binding to carbohydrate ligand.
- Additional family members are then determined based on amino acid sequence similarity. To be defined as a true CBM, it must form part of a larger amino acid sequence encoding a putative CAZyme (or enzyme with demonstrated activity on a carbohydrate-containing substrate and the CBM enhances the catalytic efficiency of the enzyme by binding with or in close proximity of the substrate).
Amino acid sequence-based classification of a CBM family may lead to the incorporation of other carbohydrate binding proteins within a given family, including lectins (such as ricin (CBM13), tachycitin (CBM14), wheat germ agglutinin (CBM18), fucolectin (CBM47), and malectin (CBM57)) and periplasmic solute binding proteins (such asCBM32). The community is open to incorporation of all carbohydrate-binding proteins within the CBM classification system based on the above criteria.
Studying CBM-ligand Interactions
Laboratory approaches to studying the binding function of carbohydrate-binding modules has been extensively reviewed by Abbott and Boraston [16]. Typically, molecular biology techniques are used to overproduce a CBM protein in a host strain such as Escherichia coli which is then isolated and purified. Initial screening for carbohydrate binding interactions can be performed using microarray techniques [17] or other screening methods (ref). Several approaches can be taken to verify and quantify CBM-polysaccharide interactions. Key approaches include fluorescence microscopy on plant or mammalian tissues [17] [18], affinity gel electrophoresis (ref), UV difference and fluorescence spectroscopy (ref), solid state depletion assay (ref) and isothermal titration calorimetry [19].
Overall demonstration of carbohydrate binding function by CBMs is essential to understanding how these associated modules confer enzymatic specificity to carbohydrate-active enzymes.
References
-
Davies, G.J. and Sinnott, M.L. (2008) Sorting the diverse: the sequence-based classifications of carbohydrate-active enzymes. Biochem. J. (BJ Classic Paper, online only). DOI: 10.1042/BJ20080382
- Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, and Henrissat B. (2009). The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res. 2009;37(Database issue):D233-8. DOI:10.1093/nar/gkn663 |
- Boraston AB, Bolam DN, Gilbert HJ, and Davies GJ. (2004). Carbohydrate-binding modules: fine-tuning polysaccharide recognition. Biochem J. 2004;382(Pt 3):769-81. DOI:10.1042/BJ20040892 |
- Hashimoto H (2006). Recent structural studies of carbohydrate-binding modules. Cell Mol Life Sci. 2006;63(24):2954-67. DOI:10.1007/s00018-006-6195-3 |
- Shoseyov O, Shani Z, and Levy I. (2006). Carbohydrate binding modules: biochemical properties and novel applications. Microbiol Mol Biol Rev. 2006;70(2):283-95. DOI:10.1128/MMBR.00028-05 |
- Guillén D, Sánchez S, and Rodríguez-Sanoja R. (2010). Carbohydrate-binding domains: multiplicity of biological roles. Appl Microbiol Biotechnol. 2010;85(5):1241-9. DOI:10.1007/s00253-009-2331-y |
- Boraston AB, Ficko-Blean E, and Healey M. (2007). Carbohydrate recognition by a large sialidase toxin from Clostridium perfringens. Biochemistry. 2007;46(40):11352-60. DOI:10.1021/bi701317g |
- Adams JJ, Gregg K, Bayer EA, Boraston AB, and Smith SP. (2008). Structural basis of Clostridium perfringens toxin complex formation. Proc Natl Acad Sci U S A. 2008;105(34):12194-9. DOI:10.1073/pnas.0803154105 |
- Lammerts van Bueren A, Ficko-Blean E, Pluvinage B, Hehemann JH, Higgins MA, Deng L, Ogunniyi AD, Stroeher UH, El Warry N, Burke RD, Czjzek M, Paton JC, Vocadlo DJ, and Boraston AB. (2011). The conformation and function of a multimodular glycogen-degrading pneumococcal virulence factor. Structure. 2011;19(5):640-51. DOI:10.1016/j.str.2011.03.001 |
- Ficko-Blean E, Gregg KJ, Adams JJ, Hehemann JH, Czjzek M, Smith SP, and Boraston AB. (2009). Portrait of an enzyme, a complete structural analysis of a multimodular {beta}-N-acetylglucosaminidase from Clostridium perfringens. J Biol Chem. 2009;284(15):9876-84. DOI:10.1074/jbc.M808954200 |
-
Van Tilbeurgh, H., Tomme P., Claeyssens M., Bhikhabhai R., Pettersson G.(1986) Limited proteolysis of the cellobiohydrolase I from Trichoderma reesei. FEBS Lett. 204,223–227. [1]
- Tomme P, Van Tilbeurgh H, Pettersson G, Van Damme J, Vandekerckhove J, Knowles J, Teeri T, and Claeyssens M. (1988). Studies of the cellulolytic system of Trichoderma reesei QM 9414. Analysis of domain function in two cellobiohydrolases by limited proteolysis. Eur J Biochem. 1988;170(3):575-81. DOI:10.1111/j.1432-1033.1988.tb13736.x |
- Gilkes NR, Warren RA, Miller RC Jr, and Kilburn DG. (1988). Precise excision of the cellulose binding domains from two Cellulomonas fimi cellulases by a homologous protease and the effect on catalysis. J Biol Chem. 1988;263(21):10401-7. | Google Books | Open Library
-
Tomme, P., Warren, R.A., Miller, R.C., Jr., Kilburn, D.G. & Gilkes, N.R. (1995) in Enzymatic Degradation of Insoluble Polysaccharides (Saddler, J.N. & Penner, M., eds.), Cellulose-binding domains: classification and properties. pp. 142-163, American Chemical Society, Washington.
- Meyer EA, Castellano RK, and Diederich F. (2003). Interactions with aromatic rings in chemical and biological recognition. Angew Chem Int Ed Engl. 2003;42(11):1210-50. DOI:10.1002/anie.200390319 |
- Abbott DW and Boraston AB. (2012). Quantitative approaches to the analysis of carbohydrate-binding module function. Methods Enzymol. 2012;510:211-31. DOI:10.1016/B978-0-12-415931-0.00011-2 |
- van Bueren AL, Higgins M, Wang D, Burke RD, and Boraston AB. (2007). Identification and structural basis of binding to host lung glycogen by streptococcal virulence factors. Nat Struct Mol Biol. 2007;14(1):76-84. DOI:10.1038/nsmb1187 |
- McCartney L, Blake AW, Flint J, Bolam DN, Boraston AB, Gilbert HJ, and Knox JP. (2006). Differential recognition of plant cell walls by microbial xylan-specific carbohydrate-binding modules. Proc Natl Acad Sci U S A. 2006;103(12):4765-70. DOI:10.1073/pnas.0508887103 |
- Lammerts van Bueren A and Boraston AB. (2004). Binding sub-site dissection of a carbohydrate-binding module reveals the contribution of entropy to oligosaccharide recognition at "non-primary" binding subsites. J Mol Biol. 2004;340(4):869-79. DOI:10.1016/j.jmb.2004.05.038 |