CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycoside Hydrolase Family 35
This page has been approved by the Responsible Curator as essentially complete. CAZypedia is a living document, so further improvement of this page is still possible. If you would like to suggest an addition or correction, please contact the page's Responsible Curator directly by e-mail.
- Authors: Anna Kulminskaya, Mirko Maksimainen, Juha Rouvinen, Masahiro Nakajima
- Responsible Curator: Anna Kulminskaya
| Glycoside Hydrolase Family GH35 | |
| Clan | GH-A |
| Mechanism | retaining |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/GH35.html | |
Substrate specificities
The majority of glycoside hydrolases of GH35 are β-galactosidases (EC 3.2.1.23). GH35 enzymes have been isolated from microorganisms such as fungi, bacteria and yeasts, as well as higher organisms such as plants, animals, and human cells. These β-galactosidases catalyse the hydrolysis of terminal non-reducing β-D-galactose residues in, for example, lactose (1,4-O-β-D-galactopyranosyl-D-glucose), oligosaccharides, glycolipids, and glycoproteins. Various GH35 β-galactosidases demonstrate specificity towards β-1,3-, β-1,6- or β-1,4-galactosidic linkages [1, 2, 3], and are often most active under acidic conditions [4, 5, 6]. As with many other CAZy families [7, 8, 9], GH35 members tend to be represented by multi-gene families in plants [3, 10, 11, 12, 13]. Moreover, plant GH35 β-galactosidases have be divided into two classes: members of the first are capable of hydrolyzing pectic β-1,4-galactans, while those of the second can specifically cleave β-1,3- and β-1,6-galactosyl linkages of arabinogalactan proteins [14]. In 2025, a β-galactosidase acting specifically on β-1,2-galactooligosaccharides (EC 3.2.1.230) was reported [15].
In addition to β-galactosidases, GH35 also contains a limited number of archeal exo-β-glucosaminidases (EC 3.2.1.165) [16, 17] and β-1,2-glucosyltransferase (EC 2.4.1.391) [18]. The former enzymes hydrolyze chitosan or chitosan oligosaccharides to remove successive D-glucosamine residues from non-reducing termini. The latter enzyme disproportionates β-1,2-glucooligosaccharides by transferring glucose units but prefers sophorose (Glc-β-1,2-Glc) as a donor and alkyl- or acyl-glucosides as acceptors, regardless of their anomeric configuration.
Kinetics and Mechanism
Beta-galactosidases of GH35 catalyze the hydrolysis of terminal β-galactosyl residues via a classical Koshland retaining mechanism, which leads to net retention of the β-anomeric configuration of the released galactose molecule. The stereochemistry of the reaction was first shown by NMR for the human β-galactosidase precursor [4] and has been subsequently confirmed by other investigators for microbial and plant enzymes [1, 5] .
Catalytic Residues
The catalytic residues for family 35 were first predicted on the basis of hydrophobic cluster analysis of proteins of similar protein fold [19]. Experimentally, the glutamic acid residue 268 was first identified as the catalytic nucleophile in human lysosomal β-galactosidase precursor using a slow substrate, 2,4-dinitrophenyl 2-deoxy-2-fluoro-β-D-galactopyranoside, which allowed trapping of a covalent glycosyl-enzyme intermediate and subsequent peptide mapping [20]. This approach was repeated for two bacterial β-galactosidases from Xanthomonas manihotis and Bacillus circulans [21]. The general acid/base residue was inferred to be Glu200 from structural studies of a Penicillium sp. β-galactosidase [22]. Recent structural studies (vide infra) revealed two different conformations of the general acid/base residue in the β-galactosidase of Trichoderma reesei [23].
Three-dimensional structures
The first 3D-structures of a GH35 enzyme, those of a β-galactosidase from Pencillium sp. (Psp-β-gal) in native (PDB 1tg7) and product-complexed (PDB 1xc6) forms, were reported in 2004 at 1.90 Å and 2.10 Å resolution, respectively [22]. The structure of a β-galactosidase from Bacteriodes thetaiotamicron (Btm-β-gal) was subsequently reported by the New York Structural GenomiX Research Consortium in 2008 at 2.15 Å resolution (PDB 3d3a). In 2010, an atomic (1.2 Å) resolution crystal structure of a Trichoderma reesei (Hypocrea jecorina) β-galactosidase (Tr-β-gal, PDB 3og2) was reported, together with complex structures with galactose, IPTG and PETG at 1.5, 1.75 and 1.4 Å resolutions, respectively (PDB codes 3ogr, 3ogs, and 3ogv, respectively) [23].
GH35 enzymes belong to Clan GH-A, and thus have an (α/β)8 (TIM) barrel as the catalytic domain, in which two glutamic acid residues act as the general acid-base and nucleophilic catalysts. These residues are located in strands 4 and 7 of the barrel.
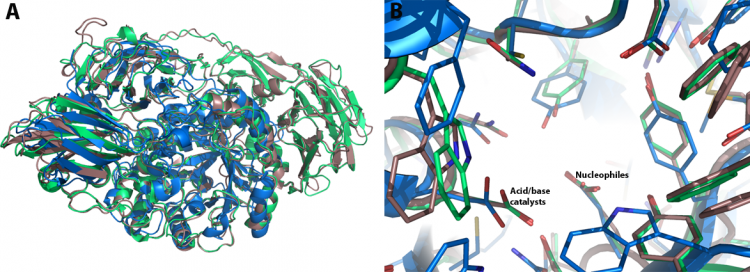
The comparison of the native structures of Psp-β-gal, Tr-β-gal and Btmβ-gal reveals two things (Figure 1): Firstly, Btm-β-gal consists of three distinct domains, whereas Psp-β-gal and Tr-β-gal consist of five and six domains, respectively. The second and third domains of Btm-β-gal are quite similar with the fourth and fifth domains of Psp-β-gal, and with the fifth and sixth domains of Tr-β-gal. Secondly, major structural differences between Psp-β-gal and Tr-β-gal are in the conformations of the loop regions. Although the crystal structures of Psp-β-gal and Tr-β-gal are similar, the interpretation of the structure of Tr-β-gal is somewhat different from that presented earlier for Psp-β-gal: Rojas et al. considered Psp-β-gal to be composed of five distinct structural domains. The overall structure is built around the first, TIM barrel, domain. Domain 2 is an all β-sheet domain containing an immunoglobulin-like subdomain, Domain 3 is based on a Greek-key β-sandwich, and Domains 4 and 5 are jelly rolls [22]. In contrast, Maksimainen et al. concluded the domain 2 includes two different domains and thus the Tr-β-gal and Psp-β-gal structures both form six similar domains [23].
The superimposition of the active sites of the GH35 β-galactosidases shows a remarkable similarity. In addition to the catalytic residues, the active sites of the GH35 β-galactosidases contain many identical residues (Figure 1B). Based on the galactose-bound crystallographic models of Psp-β-gal and Tr-β-gal, a single galactose molecule is bound to the active site of the GH35 enzyme in the chair conformation in the β-anomeric configuration.
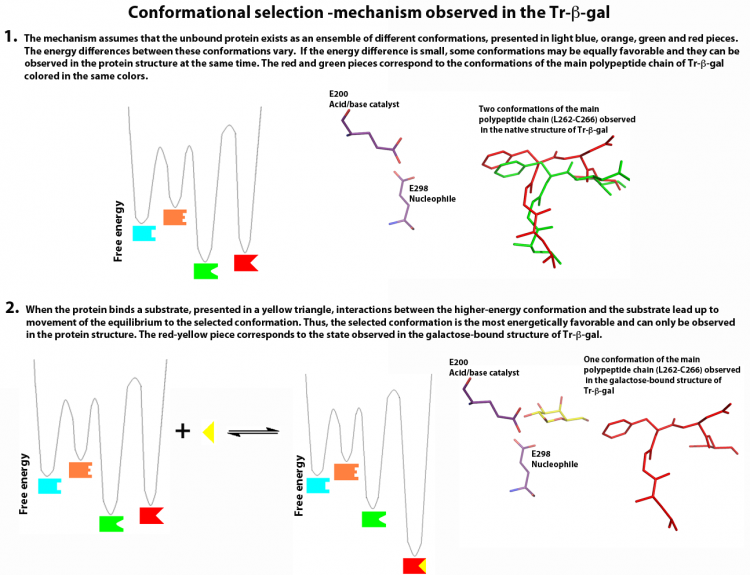
Additionally, Maksimainen et al. have described conformational changes in two loop regions of the active site of Tr-β-gal, that implicates a conformational selection mechanism for the enzyme (Figure 2). Unlike the induced fit theory, which assumes that the initial interaction between a protein and its binding partner induces a conformational change in the protein through a stepwise process, the conformational selection theory is based on the assumption that the unbound protein exists as an ensemble of conformations in dynamic equilibrium. Interaction between a weakly populated, higher-energy conformation and a binding partner causes the equilibrium to move in favor of the selected conformation [24, 25]. This can be seen in the structures of Tr-β-gal: the open and closed conformation are both favorable in the native structure and the closed conformation becomes more favorable in the complex structures. Furthermore, The acid/base catalyst Glu200 has two different conformations in the IPTG and PETG complex structures that clearly affects the pKa value of this residue and thus the catalytic mechanism of the enzyme [23].
Structure images
Family Firsts
- First stereochemistry determination
Retaining stereochemical outcome for human β-galactosidase precursor by NMR [4]
- First catalytic nucleophile identification
Human β-galactosidase precursor by 2-fluorogalactose labeling [20].
- First general acid/base residue identification
Penicillium sp. β-galactosidase by structural identification [22].
- First 3-D structure
Penicillium sp. β-galactosidase [22].
References
- Zinin AI, Eneyskaya EV, Shabalin KA, Kulminskaya AA, Shishlyannikov SM, and Neustroev KN. (2002). 1-O-Acetyl-beta-D-galactopyranose: a novel substrate for the transglycosylation reaction catalyzed by the beta-galactosidase from Penicillium sp. Carbohydr Res. 2002;337(7):635-42. DOI:10.1016/s0008-6215(02)00027-7 |
- Gamauf C, Marchetti M, Kallio J, Puranen T, Vehmaanperä J, Allmaier G, Kubicek CP, and Seiboth B. (2007). Characterization of the bga1-encoded glycoside hydrolase family 35 beta-galactosidase of Hypocrea jecorina with galacto-beta-D-galactanase activity. FEBS J. 2007;274(7):1691-700. DOI:10.1111/j.1742-4658.2007.05714.x |
- Tanthanuch W, Chantarangsee M, Maneesan J, and Ketudat-Cairns J. (2008). Genomic and expression analysis of glycosyl hydrolase family 35 genes from rice (Oryza sativa L.). BMC Plant Biol. 2008;8:84. DOI:10.1186/1471-2229-8-84 |
- Zhang S, McCarter JD, Okamura-Oho Y, Yaghi F, Hinek A, Withers SG, and Callahan JW. (1994). Kinetic mechanism and characterization of human beta-galactosidase precursor secreted by permanently transfected Chinese hamster ovary cells. Biochem J. 1994;304 ( Pt 1)(Pt 1):281-8. DOI:10.1042/bj3040281 |
- van Casteren WH, Eimermann M, van den Broek LA, Vincken JP, Schols HA, and Voragen AG. (2000). Purification and characterisation of a beta-galactosidase from Aspergillus aculeatus with activity towards (modified) exopolysaccharides from Lactococcus lactis subsp. cremoris B39 and B891. Carbohydr Res. 2000;329(1):75-85. DOI:10.1016/s0008-6215(00)00152-x |
- Wang H, Luo H, Bai Y, Wang Y, Yang P, Shi P, Zhang W, Fan Y, and Yao B. (2009). An acidophilic beta-galactosidase from Bispora sp. MEY-1 with high lactose hydrolytic activity under simulated gastric conditions. J Agric Food Chem. 2009;57(12):5535-41. DOI:10.1021/jf900369e |
- Geisler-Lee J, Geisler M, Coutinho PM, Segerman B, Nishikubo N, Takahashi J, Aspeborg H, Djerbi S, Master E, Andersson-Gunnerås S, Sundberg B, Karpinski S, Teeri TT, Kleczkowski LA, Henrissat B, and Mellerowicz EJ. (2006). Poplar carbohydrate-active enzymes. Gene identification and expression analyses. Plant Physiol. 2006;140(3):946-62. DOI:10.1104/pp.105.072652 |
- Henrissat B, Coutinho PM, and Davies GJ. (2001). A census of carbohydrate-active enzymes in the genome of Arabidopsis thaliana. Plant Mol Biol. 2001;47(1-2):55-72. | Google Books | Open Library
- Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, Schein J, Sterck L, Aerts A, Bhalerao RR, Bhalerao RP, Blaudez D, Boerjan W, Brun A, Brunner A, Busov V, Campbell M, Carlson J, Chalot M, Chapman J, Chen GL, Cooper D, Coutinho PM, Couturier J, Covert S, Cronk Q, Cunningham R, Davis J, Degroeve S, Déjardin A, Depamphilis C, Detter J, Dirks B, Dubchak I, Duplessis S, Ehlting J, Ellis B, Gendler K, Goodstein D, Gribskov M, Grimwood J, Groover A, Gunter L, Hamberger B, Heinze B, Helariutta Y, Henrissat B, Holligan D, Holt R, Huang W, Islam-Faridi N, Jones S, Jones-Rhoades M, Jorgensen R, Joshi C, Kangasjärvi J, Karlsson J, Kelleher C, Kirkpatrick R, Kirst M, Kohler A, Kalluri U, Larimer F, Leebens-Mack J, Leplé JC, Locascio P, Lou Y, Lucas S, Martin F, Montanini B, Napoli C, Nelson DR, Nelson C, Nieminen K, Nilsson O, Pereda V, Peter G, Philippe R, Pilate G, Poliakov A, Razumovskaya J, Richardson P, Rinaldi C, Ritland K, Rouzé P, Ryaboy D, Schmutz J, Schrader J, Segerman B, Shin H, Siddiqui A, Sterky F, Terry A, Tsai CJ, Uberbacher E, Unneberg P, Vahala J, Wall K, Wessler S, Yang G, Yin T, Douglas C, Marra M, Sandberg G, Van de Peer Y, and Rokhsar D. (2006). The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science. 2006;313(5793):1596-604. DOI:10.1126/science.1128691 |
- Ahn YO, Zheng M, Bevan DR, Esen A, Shiu SH, Benson J, Peng HP, Miller JT, Cheng CL, Poulton JE, and Shih MC. (2007). Functional genomic analysis of Arabidopsis thaliana glycoside hydrolase family 35. Phytochemistry. 2007;68(11):1510-20. DOI:10.1016/j.phytochem.2007.03.021 |
- Smith DL and Gross KC. (2000). A family of at least seven beta-galactosidase genes is expressed during tomato fruit development. Plant Physiol. 2000;123(3):1173-83. DOI:10.1104/pp.123.3.1173 |
- Lazan H, Ng SY, Goh LY, and Ali ZM. (2004). Papaya beta-galactosidase/galactanase isoforms in differential cell wall hydrolysis and fruit softening during ripening. Plant Physiol Biochem. 2004;42(11):847-53. DOI:10.1016/j.plaphy.2004.10.007 |
- Ross GS, Wegrzyn T, MacRae EA, and Redgwell RJ. (1994). Apple beta-galactosidase. Activity against cell wall polysaccharides and characterization of a related cDNA clone. Plant Physiol. 1994;106(2):521-8. DOI:10.1104/pp.106.2.521 |
- Kotake T, Dina S, Konishi T, Kaneko S, Igarashi K, Samejima M, Watanabe Y, Kimura K, and Tsumuraya Y. (2005). Molecular cloning of a {beta}-galactosidase from radish that specifically hydrolyzes {beta}-(1->3)- and {beta}-(1->6)-galactosyl residues of Arabinogalactan protein. Plant Physiol. 2005;138(3):1563-76. DOI:10.1104/pp.105.062562 |
- Nakazawa Y, Kageyama M, Matsuzawa T, Liang Z, Kobayashi K, Shimizu H, Maeda K, Masuhiro M, Motouchi S, Kumano S, Tanaka N, Kuramochi K, Nakai H, Taguchi H, and Nakajima M. (2025). Structure and function of a β-1,2-galactosidase from Bacteroides xylanisolvens, an intestinal bacterium. Commun Biol. 2025;8(1):66. DOI:10.1038/s42003-025-07494-1 |
- Liu B, Li Z, Hong Y, Ni J, Sheng D, and Shen Y. (2006). Cloning, expression and characterization of a thermostable exo-beta-D-glucosaminidase from the hyperthermophilic archaeon Pyrococcus horikoshii. Biotechnol Lett. 2006;28(20):1655-60. DOI:10.1007/s10529-006-9137-0 |
- Kobayashi K, Shimizu H, Tanaka N, Kuramochi K, Nakai H, Nakajima M, and Taguchi H. (2022). Characterization and structural analyses of a novel glycosyltransferase acting on the β-1,2-glucosidic linkages. J Biol Chem. 2022;298(3):101606. DOI:10.1016/j.jbc.2022.101606 |
- Henrissat B, Callebaut I, Fabrega S, Lehn P, Mornon JP, and Davies G. (1995). Conserved catalytic machinery and the prediction of a common fold for several families of glycosyl hydrolases. Proc Natl Acad Sci U S A. 1995;92(15):7090-4. DOI:10.1073/pnas.92.15.7090 |
- McCarter JD, Burgoyne DL, Miao S, Zhang S, Callahan JW, and Withers SG. (1997). Identification of Glu-268 as the catalytic nucleophile of human lysosomal beta-galactosidase precursor by mass spectrometry. J Biol Chem. 1997;272(1):396-400. DOI:10.1074/jbc.272.1.396 |
- Blanchard JE, Gal L, He S, Foisy J, Warren RA, and Withers SG. (2001). The identification of the catalytic nucleophiles of two beta-galactosidases from glycoside hydrolase family 35. Carbohydr Res. 2001;333(1):7-17. DOI:10.1016/s0008-6215(01)00108-2 |
- Rojas AL, Nagem RA, Neustroev KN, Arand M, Adamska M, Eneyskaya EV, Kulminskaya AA, Garratt RC, Golubev AM, and Polikarpov I. (2004). Crystal structures of beta-galactosidase from Penicillium sp. and its complex with galactose. J Mol Biol. 2004;343(5):1281-92. DOI:10.1016/j.jmb.2004.09.012 |
- Maksimainen M, Hakulinen N, Kallio JM, Timoharju T, Turunen O, and Rouvinen J. (2011). Crystal structures of Trichoderma reesei β-galactosidase reveal conformational changes in the active site. J Struct Biol. 2011;174(1):156-63. DOI:10.1016/j.jsb.2010.11.024 |
- Tsai CJ, Ma B, and Nussinov R. (1999). Folding and binding cascades: shifts in energy landscapes. Proc Natl Acad Sci U S A. 1999;96(18):9970-2. DOI:10.1073/pnas.96.18.9970 |
- Boehr DD and Wright PE. (2008). Biochemistry. How do proteins interact?. Science. 2008;320(5882):1429-30. DOI:10.1126/science.1158818 |
- Tanaka T, Fukui T, Atomi H, and Imanaka T. (2003). Characterization of an exo-beta-D-glucosaminidase involved in a novel chitinolytic pathway from the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J Bacteriol. 2003;185(17):5175-81. DOI:10.1128/JB.185.17.5175-5181.2003 |